Introduction
Plants in nature are always threatened by different biotic and abiotic stresses among which biotic stress caused by different pathogens plays an important role and sometimes causes upto 20% yield loss in production of crops in worldwide1. With the passing time and advances in research, more and more active compounds have been developed which can control the pathogen attack, but still they are not fully successful as because there are different challenges like resistance against chemicals, mutation controlled pathogen supremacy, environmental hazard, health concern, effect on non target organism etc. For identifying the solution of these burning issues, the crop disease management through sustainable methods has come upon the rows.According to Cook and Baker (1983), “Biological control is the reduction of the amount of inoculum or diseaseproducing activity of a pathogen accomplished by or through one or more organisms other than man”2. Basically biological control not only depends upon predatism and parasitism but also on antibiosis, induced defense mechanism, competitive suppressiveness and the production of defense related chemicals like enzymes, phytolexin, toxin etc.3-5 Enhancement of the plant innate immunity and strengthening of defense mechanism by acquired and induced way by the activity of other organism is becoming a new line of thought for the researchers. This type of beneficial microorganisms are generally centralized in the rhizosphere of the plant.6 By the natural activity, plants are blessed with the secretion of different complex compounds like phenols, propanoids, sugars in the root zone which somehow harbors these kind of microorganisms and turns the rhizosphere into an amazing place with the enriched microbes diversity and biomass. In this review article we will focus on the connection between these three components i.e. root exudates, rhizosphere biodiversity and their effect on plant defense mechanism. Their well-established interaction, mass effect and the perspective of the trident will be discussed here in elaborative manner.
Root exudates-A shield of plant base
Root exudates are the complex organic substances that are secreted by plant through its different physiological activity and freely releases root-cell material or mucilage and deposits in rhizosphere zone.7It is estimated that at least 40% carbon released from photosynthetic process released as different exudates through plant roots.8,9 The secretion of the plant as root exudates greatly influenced by different factors like environmental stimulation,10 physiological condition of plants, plant species11 and development stages.12 The exudates from roots are generally classified into two broad categories i.e. low molecular amino acids, organic acids, sugars and other secondary metabolites (having low molecular weight) and mucilage (polysaccharides), proteins etc. (high molecular weight).13 It is also not a true fact that root exudates should be very unique to a plant as the same can be secreted by different plant species and even in various life stages e.g. benzoxazinoids (BXs) found to be produced by different species of poaceae family including maize, wheat, and rye.14 The chemicals found in rhizosphere zone are not always secreted by plant itself but sometimes they can be produce by cell lysis whichis mediated by cell rupturing due to different biotic and abiotic damage.7
Table 1: Root exudates profiles of different crops.
|
Sl.no. |
Associated Plant |
Root exudates |
Chemical Nature of root exudates |
Growth Media/ Substrate |
Reference |
|
1 |
Tomato |
Citric Acid, Piruvic Acid, Malic Acid, Succinic Acid, Fumaric, Pyroglutamic |
Organic Acid |
Stone wool and Glass beads |
15 |
|
Glucose, Fructose, Maltose, Ribose, Xylose |
Sugars |
||||
|
Tryptophane |
Growth hormone |
||||
|
2 |
Cucumber |
Citric Acid, Piruvic Acid, Malic Acid, Succinic Acid, Fumaric, Pyroglutamic |
Organic Acid |
Stone wool and Glass beads |
16 |
|
Glucose, Fructose, Maltose, Ribose, Xylose, Melibiose |
Sugars |
||||
|
Tryptophane |
Growth hormone |
||||
|
3 |
Lettuce |
Alanine,Aspertate,Glutamine,Glysine, Leucine,Isoleucine, Proline,Serine,Threonine,Valine,Prutresine |
Amino acids and amines |
Loess Loam,Alluvial Loam |
17 |
|
Fructose,Maltose,Trehalose,Sucrose,Glycerol |
Sugars and Sugar alcohols |
||||
|
Malate,Fumarate,Succinate, Lauric Acid,Benzoic Acid |
Organic Acid |
||||
|
4. |
Banana |
Oxalic, Malic and Fumaric acid |
Organic Acid |
Vermiculite |
18 |
|
5. |
Water melon |
Gallic acid, Coumaric acid, β-hydroxybenzoic acid, Chlorogenic acid, Vanillic acid, Caffeic acid, Syringic acid, Ferulic acid, Benzoic acid, Salicylic acid and Cinnamic acid |
Phenolic Compund |
Mixture of surface soil |
19 |
|
6. |
Cotton |
Glucose,Xylose |
Sugars |
Mixture of soil |
20 |
|
Lysin,Threonine, Arginine |
Amino acids |
||||
|
Oxalic acid,Succinic Acid,Maleic Acid |
Organic Acid |
||||
|
7. |
Wheat |
Glucose,Fructose,Sucrose, Xylose |
Sugars |
Soil mixture |
20 |
|
|
|
||||
|
Arginine,Threonine, Methionine |
Amino Acids |
||||
|
|
|
Succinic, Oxalic,Malonic |
Organic Acid |
|
|
|
|
Relationship of root exudates and soil microbiomes
The plants are well known for engaging different beneficial microbiomes that actively do their assigned activity and this incidence have ample evidences that cannot be denied.21 The chemical cross talk between the microbiomes and host plant make the primary basis and foundation bridge among themselves (Fig.1). Additionally, it demonstrated how the host plant actually prepares and blueprints its shape, bulk and affinity of their microbiomes in root zones by the presence, composition and quantity of the exudation which directly or indirectly but actively impact on soil invading or soil inhibiting micro-organisms.22 Before illustrating the mode of action in between the primary metabolite of root exudation and microbiomes’ reaction towards them, we have to concentrate on the most sensitive part of plants root i.e. root tip which is the first to react or expose into the environmental stimulation.23 Rhizosphere is also well known as microbial hot spot of the rhizosphere24 where bacterial population scored highest followed by other microorganism like fungi, algae and actinomycetes25 The thumb rule for the successful communication between the host plant and microbes solely depends on the two main factors; the production and circulation of specified signals and the responses produce by this signals taking as stimuli26 In the process of symbiosis, the Nod-factors or lipo-chitooligosaccharides that act as the signaling molecule of nodulation generally get activated by the secondary metabolite like flavonoids.27 These Nod-factors actually plays an important role by triggering plant growth leading to formation of root nodule through the changes in root hair structure.On the other hand, it was also proven that presence of micro-organism in plant growth solution actually enhance the root exudation. e.g. secretion from the cell wall of Phytophthora cinnamoni mediate secretion of rosmarinic acid as root exudate in the rhizosphere of sweet basil.13 In case of the antagonistic relationship between the beneficial microbiomes and soil borne pathogen, colonization in the rhizosphere is the first and foremost step between them. According to the popular “rhizosphere effect” described by Hiltner (1904) nutrients exuded by plants roots are the key attraction for the vicinity of the microorganism in the root zone. In this condition a cross talk generally ignited where carbon-rich environment produced by plant root through exudation produce signals for microbiome and in reply responses for colonization produce by microbiome. The most crucial characteristic of the microbiomes is motility, which determines the kind of relationship between the pathogenic and beneficial bacteria and which of them will be successful in colonizing the environment through this chemotaxis process.6 This chemotaxis process regulates the maximum colonization process in rhizosphere whereas some of the portion, also involved electro genic process. As an illustration, consider the application of electro physical potential in plant roots, which convey at the root surface and draw swimming oomycete plant disease zoospores to plant rhizosphere.13 From one of the compelling theory for root exudation describe pressure-driven mechanism of Munch’s phloem flow, responsible for the transport of carbon in source organs to sinks through differences in concentration gradients that vary in turgor pressure based on source-sink activities, controls the driving carbon transport to roots and then extraction from roots to soil.28, 29 In this regard, the statement of Ross-Elliott et al. (2017) described that in the root tip portion, phloem/batch unloading occurs through plasmodesmata by the combination of mass flow and diffusion.30
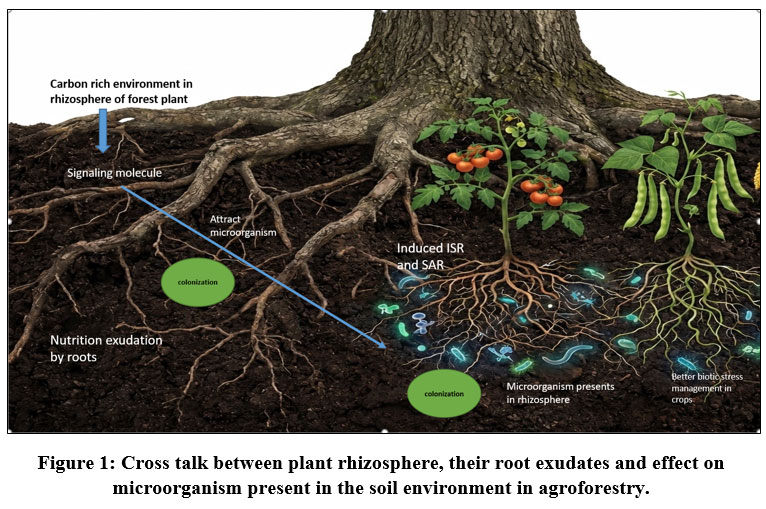 |
Figure 1: Cross talk between plant rhizosphere, their root exudates and effect on microorganism present in the soil environment in agroforestry.
|
Transportation of Primary Metabolites in extra cellular zone of root tip
As mentioned previously, plasma membrane considered as the permeable to gas and molecules while impermeable to charged molecules like ions. As a result, these molecules can only pass through particular transmembrane proteins that create tiny holes in the bilayer lipid zone, enabling charged molecules to go through the membrane excluding the chances of being in contact with the fatty acid chains of the phospholipids which is hydrophobic in nature.23 Many researchers mentioned about the efflux carriers readily channels present to mediate the exudation flux regulation controlled by gene expression.
A list of metabolites has been enlisted below with their efflux transporters
Table 2: Primary metabolites and corresponding transporters
|
Sl. No. |
Primary metabolites |
Transporters |
Reference |
|
1 |
Amino acids |
UMAMIT transporters |
31 |
|
|
|
CAT transporters |
32 |
|
|
|
GDU transporters |
33 |
|
2 |
Sugars |
SWEET transporter |
34 |
|
3 |
Organic acids |
ALMT/malate and MATE/citrate transporters |
35 |
In some of cases, nutrient imbalances, environmental factors or some edaphic factors are also seemed to be responsible to regulate the gene transcription related to specific efflux transporter. For an instance, expression of SWEET, activated by the pathogen-driven molecule increase glucose efflux into the root apoplast36 whereas, some metallic ion like Al3+toxicity or scarcity of P triggers the secretion of the malate by the process of ALMT up-regulation mechanism in rhizosphere.35 After removal of metabolites from cell plasma membrane of phloem used by the microbes of the rhizosphere. After attaining the apoplastic pathway, the secondary work is the well-defined diffusion process in the mentioned zone lacking of no barrier produced by the plasma membranes through root tip. Diffusion rate also differs from each other according to the zone of root tip. As an instance, in the immature elongation zone it is found to be higher rate of diffusion up to 10 times than the mature zone of root tip in the roots of Arabidopsis.37
Networking system in rhizosphere
The microorganism residing in the rhizosphere zone generally act as fostering community that community that participate in plant development and plant health.37 This rich microorganism biodiversity and its composition vary with the two main factors, i.e.plant species and soil properties.38 Among the same species the assembly of rhizobacterial communities shows the differences even with the differences of genotypes and on the other hand can be same even in different environment and in different region.39,40 Plants are generally select the microbiome network in their rhizosphere according to their needs for fitness like particular need of nutrient of plants, biotic stress control procedure and symbiotic relationship etc.41 As an example, root inhibiting bacteria were generally found to be well established in the grassland of C3 grasses than the grassland of C4 grasses on the other hand, Burkholderia and Variovorax were found enriched in the rhizosphere of Leguminosae, while the root zone of Gramineaepredominated by Pedobacter and Aeromicrobium.42 Here some of the examples have been cited which indicates the special microbiome present in the special agroforestry cropping model.
Table 3: Microbiome profiles of forest plants and their associated agricultural crops.
|
Forest plant |
Agricultural crop |
Presence of micro-organism |
reference |
|
Poplar |
Barley and Maize |
Bradyrhizobium and Mesorhizobium |
43,44 |
|
Olive tree |
Durum wheat, Barley, Chickpea and Faba bean |
Actinobacteria , Proteobacteria , Crenarchaeota , Acidobacteria , Bacteroidetes , Planctomycetes , Firmicutes , Chloroflexi and Verrucomicrobia |
45 |
|
Poplar, willow |
Wheat, Maize, and Rapeseed |
Acidobacteria, Actinobacteria, Alpha- and Gammaproteobacteria, Firmicutes, and Verrucomicrobia |
46 |
|
Atractylodes lancea |
Pea nut |
Gram-negative (G−) bacteria |
47 |
Plant defense modulators
Root exudates secreted in the rhizosphere zone not only helpful for modifying the soil’s own physiochemical property but also modulate the proportion of microbial communities which is directly or indirectly helpful for the bioremediation of soil contaminants and as well as plant defense against the pathogen (Fig.1). This defense mechanism centered in creating the endophytic mutualism, development of antimicrobial activity and mimicry of quorum sensing etc.48 Although Vicré et al. (2005) has described a different defense mechanism called border cell organization in Arabidopsis, where specialized root cells with the capability of active secretion of metabolites, would be arranged not only to attract the beneficial microorganisms but to engulf harmful disease causing prokaryotes and nematode by producing mucilage covering.49 The mechanism and the specificity of targeting pathogen differ from microbes to microbe and depends on even root exudates variation. On the other hand Pseudomonas spp. Specific traits for colonization in a particular rhizosphere zone depends on the flagella motility which mediated by the amount of secretion in root zone.50 Here the interesting fact is that sometimes the presence of some specific microflaura resulted in the changes of composition in rhizosphere zone.As an example use of biocontrol strain Pseudomonas fluorescens WCS365 (WCS365) on tomato roots increase the quantity of the total organic acids.15 The augmentation of antagonistic bacteria of fluorescent Pseudomonas spp strain showed a certain decline Gaeumannomyces graminis var. tritici (Ggt) causing take all disease in wheat monoculture field where the pathogen accumulation was very severe due to repetitive cultivation. This take-all decline (TAD) wheat rhizosphere was due secretion ofantibiotic 2,4-diacetylphloroglucinol (DAPG)by antagonist51 (Kwak et al., 2009). Long chain fatty acids and amino acids secretion in rhizosphere zone proliferate the presence of Sphingomonas, Pseudomonas, Roseiflexus, and Flavitalea helpful for suppressing the foliar pathogen.52 Pseudomonas syringae pv. tomato DC3000 (Pst) causing the leaf infection in Arabidopsis encouraged the secretion of large amounts of malic acid acted receptor of chemical sensitivityfor the quorum sensing of Bacillus subtilisFB17 in the rhizosphere.53 Ocimum basilicum infected with the Pythium ultimum encouraged to produce caffeic acid derivative-rosmarinic acid, which can inhibit the multiple soilborne and soil inhibiting microorganisms.13 Use of fluorescent Pseudomonas spp. has also been identified to suppress Fusarium wilt in radish54 by its tremendous root-colonizing capability and enormous production of antimicrobial compounds.55 On the other hand as a part of creating compettion for food and space the chemo attractant bioagents employes a good role.As an example, WCS358 strain of fluorescent Pseudomonas spp produce the siderophore pseudobactin-358 which makes the iron status unavailable by chealting Fe in the rhizosphere and as they have the highly specific receptor for the Fe absorption.This monopoly condition forced the pathogen to show back off as competition for iron defeat them by compromising with their growth suppression and reduced colonization.56 Pseudomonas cepacia is well known for its capability of the production of lytic enzymes are such as cellulases, glucanases, proteases, and chitinases which suppress the Rhizoctonia solani, Sclerotium rolfsii, and P. ultimum by degrading the fungus cell structure.57
SAR and ISR in rhizosphere mediated by microbiome
Induced systemic Resistance (ISR) has been reported to be modulated by the rhizosphere bacteria although it depends on the successful interaction of the rhizosphere inhibiting strain and with the host plant species. This complex compatibility between these two depends on some environmental conditions also. Some example hence been provided for better understanding:
fluorescens WCS417r has been reported to be evoked the ISR in radish where the unsuccessful WCS358r strain in case of radish was found to be effective to elicit the ISR in Arabidopsis accession Columbia (Col-0).6
But the same WCS417r-elicited ISR reaction was failed to show ISR in case ofthe accessions RLD1 and Wassilewskija (WS-0) of Arabidopsis.58
The most important and first line of defense against any pathogen is the recognition where plant phyllo plane possess the Pattern Recognition Receptors (PRRs) which act as the recognizer of the potential harmful pathogen by working on pathogen-associated molecular patterns (PAMPs). As like as potential pathogen, the non-pathogenic microorganism produce the elicitor named Microbe-Associated Molecular Patterns (MAMPs).59 Plant immunity can be enhanced through the activation of a defensive signaling cascade triggered by the recognition of any of these PAMPs/MAMPs.6 Chinchilla et al. (2006) mentioned such an example where in Arabidopsis, the transmembrane leucine-rich-repeat receptor kinase FLS2 is directly bound by flg22, a conserved 22-amino-acid peptide of bacterial flagellin was found to activate mitogen-activated protein kinase (MAPK) signaling cascade that causes the stimulation of plant defenses by forming the antimicrobial compounds.60 Not only the microbial compounds, Plant defense are also mediated by the plant hormone secretion like Salicylic acid (SA), Jasmonic acid (JA), Abscisic acid, auxins, gibberellins, cytokinins etc. Typically, JA/ET-dependent reactions are effective against microorganism that possess necrotrophic lifestyle, while SA-dependent resistance that effective against bio trophic pathogens. SAR is related to localized deposition SA and the regulation of pathogenesis-related (PR) genes whereas, ISR regulate the JA and ET.
Conclusion
In this chapter we tried to enlighten the interconnection between plant rhizosphere and its dynamic microflora which are deviated or controlled by the root exudates. On the other hand, we discussed about the counteraction of the plant host against the harmful pathogen attacking them which is the contribution of the microflora that inhibited in rhizosphere. In this vast world of microbiomes, the identification followed by harbor the growth of beneficial microorganism and using them against the harmful one is the new tactics which can be proved as the time of need.
Article highlights
Detail idea about the rhizosphere microorganism biodiversity in different crop ecology
Detail idea about the root exudates present in the rhizosphere zone
Interaction between the micro-organism and root exudates of rhizosphere
Role of the mentioned interaction in strengthening the plant defense mechanism
Sustainable management of plant disease
Acknowledgement
We acknowledge the Dean, Faculty of Agriculture, Sri Sri University for inspiring us to contribute in this field.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of interest
The authors do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Authors’ Contributions
Debanjana Debnath: Conceptualization, Methodology, Writing – Original Draft.
Rini Labanya: Supervision, Writing – Review & Editing.
Chinmayee mohapatra: Visualization, Supervision
Suraj Goldar: Funding Acquisition, Resources.
References:
- Schumann GL, D’Arcy CJ. Essential Plant Pathology. St. Paul, MN: The American Phytopathological Society; 2006.
- Cook RJ, Baker KF. The Nature and Practice of Biological Control of Plant Pathogens. St. Paul, MN: APS Press; 1983
- Raaijmakers JM, Vlami M, de Souza JT. Antibiotic production by bacterial biocontrol agents. Antonie Van Leeuwenhoek. 2002;81:537-547. doi:10.1023/A:1020501420831
CrossRef - Haas D, Defago G. Biological control of soil-borne pathogens by fluorescent pseudomonads. Nat Rev Microbiol. 2005;3:307-319. doi:10.1038/nrmicro1129
CrossRef - Van Loon LC, Rep M, Pieterse CMJ. Significance of inducible defense-related proteins in infected plants. Annu Rev Phytopathol. 2006;44:135-162.
CrossRef - Van Wees SCM, Van der Ent S, Pieterse CMJ. Plant immune responses triggered by beneficial microbes. Curr Opin Plant Biol. 2008;11:443-448.
CrossRef - Koo BJ, Adriano DC, Bolan NS, Barton CD. Root exudates and microorganisms. In: Encyclopedia of Soils in the Environment. Vol 4. 2004:421-428
CrossRef - Curl EA, Truelove B. The Rhizosphere. Berlin: Springer-Verlag; 1986.
CrossRef - Badri DV, Vivanco JM. Regulation and function of root exudates. Plant Cell Environ. 2009;32:666-681. doi:10.1111/j.1365-3040.2009.01926.x
CrossRef - Phillips DA, Fox TC, King MD, Bhuvaneswari TV, Teuber LR. Microbial products trigger amino acid exudation from plant roots. Plant Physiol. 2004;136:2887-2894. doi:10.1104/pp.104.044222
CrossRef - Mönchgesang S, Strehmel N, Schmidt S, et al. Natural variation of root exudates in Arabidopsis thaliana: linking metabolomic and genomic data. Sci Rep. 2016;6:29033.
CrossRef - Chaparro JM, Badri DV, Vivanco JM. Rhizosphere microbiome assemblage is affected by plant development. ISME J. 2014;8:790-803.
CrossRef - Bais HP, Weir TL, Perry LG, Gilroy S, Vivanco JM. The role of root exudates in rhizosphere interactions with plants and other organisms. Annu Rev Plant Biol. 2006;57:233-266.
CrossRef - Frey M, Schullehner K, Dick R, Fiesselmann A, Gierl A. Benzoxazinoid biosynthesis: a model for evolution of secondary metabolic pathways in plants. Phytochemistry. 2009;70:1645-1651.
CrossRef - Kamilova F, Kravchenko LV, Shaposhnikov AI, Makarova N, Lugtenberg B. Effects of tomato pathogen Fusarium oxysporum f. sp. radicis-lycopersici and biocontrol bacterium Pseudomonas fluorescens WCS365 on tomato root exudates. Mol Plant Microbe Interact. 2006;19:1121-1126. doi:10.1094/MPMI-19-1121
CrossRef - Kamilova F, Kravchenko LV, Shaposhnikov AI, et al. Organic acids, sugars, and L-tryptophan in exudates of vegetables grown on stonewool and their effect on rhizosphere bacteria. Mol Plant Microbe Interact. 2006;19(3):250-256. doi:10.1094/MPMI-19-0250
CrossRef - Newman MA, Dow JM, Molinaro A, Parrilli M. Priming and modulation of plant defence responses by bacterial lipopolysaccharides. J Endotoxin Res. 2007;13:69-84.
CrossRef - Yuan J, Zhang N, Huang Q, et al. Organic acids from banana root exudates promote colonization by Bacillus amyloliquefaciens NJN-6. Sci Rep. 2015;5:13438. doi:10.1038/srep13438
CrossRef - Ling N, Zhang W, Wang D, et al. Root exudates from grafted watermelon inhibit Fusarium oxysporum f. sp. niveum. PLoS One. 2013;8(5):e63383. doi:10.1371/journal.pone.0063383
CrossRef - Narula N, Kothe E, Behl RK. Role of root exudates in plant-microbe interactions. J Appl Bot Food Qual. 2012;82:122-130.
- Reinhold-Hurek B, Bunger W, Burbano CS, Sabale M, Hurek T. Roots shaping their microbiome: hotspots for microbial activity. Annu Rev Phytopathol. 2015;53:403-424. doi:10.1146/annurev-phyto-082712-102342
CrossRef - Zolla G, Bakker MG, Badri DV, et al. Understanding root-microbiome interactions. In: De Bruijn FJ, ed. Molecular Microbial Ecology of the Rhizosphere. Hoboken, NJ: Wiley; 2013:745-754.
CrossRef - Sasse J, Martinoia E, Northen T. Feed your friends: do plant exudates shape the root microbiome? Trends Plant Sci. 2018;23:25-41.
CrossRef - Hartmann A, Rothballer M, Schmid M, Lorenz H. A pioneer in rhizosphere microbial ecology. Plant Soil. 2008;312:7-14. doi:10.1007/s11104-007-9514-z
CrossRef - Yadav BK, Akhtar MS, Panwar J. Rhizospheric plant–microbe interactions: key factors to soil fertility and plant nutrition. In: Arora NK, ed. Plant Microbes Symbiosis: Applied Facts. Springer; 2015:127-145.
CrossRef - Keller L, Surette MG. Communication in bacteria: ecological and evolutionary perspective. Nat Rev Microbiol. 2006;4:249-258. doi:10.1038/nrmicro1383
CrossRef - Spaink HP, Sheeley DM, Van Brussel AAN, et al. A novel fatty-acid moiety of lipo-oligosaccharide signals determines host specificity of rhizobium. Nature. 1991;354:125-130. doi:10.1038/354125a0
CrossRef - Münch E. Die Stoffbewegungen in der Pflanze. Jena: Carl Fischer; 1930.
CrossRef - De Schepper V, De Swaef T, Bauweraerts I, Steppe K. Phloem transport: mechanisms and controls. J Exp Bot. 2013;64:4839-4850. doi:10.1093/jxb/ert302
CrossRef - Ross-Elliott T. J., Jensen K. H., Haaning K. S., Wager B. M., Knoblauch J., Howell A. H., Phloem unloading in Arabidopsis roots is convective and regulated by the phloem-pole pericycle. elife6:2017. e24125. doi: 10.7554/eLife.24125
CrossRef - Besnard J, Zhao C, Avice JC, et al. Arabidopsis UMAMIT24 and 25 are amino acid exporters involved in seed loading. J Exp Bot. 2018;69(21):5221-5232. doi:10.1093/jxb/ery302
CrossRef - Yang, J.W., Kloepper, J.W., Ryu, C.M., 2008. Rhizosphere bacteria help plants tolerate abiotic stress. Trends Plant Sci. 14, 1–4.
CrossRef - Pratelli R, Pilot G, Regulation of amino acid metabolic enzymes and transporters in plants, Journal of Experimental Botany, Volume 65, Issue 19, October 2014, Pages 5535–5556, https://doi.org/10.1093/ jxb/eru320
CrossRef - Chen LQ, Cheung LS, Feng L, Tanner W, Frommer WB. Transport of sugars. Annu Rev Biochem. 2015;84:865-894. doi:10.1146/annurev-biochem-060614-033904
CrossRef - Mora-Macías J, Ojeda-Rivera JO, Gutiérrez-Alanís D, Yong-Villalobos L, Oropeza-Aburto A, Raya-González J, Jiménez-Domínguez G, Chávez-Calvillo G, Rellán-Álvarez R, Herrera-Estrella L.2017. Malate-dependent Fe accumulation is a critical checkpoint in the root developmental response to low phosphate. Proceedings of the National Academy of Sciences, USA 114, E3563–E3572.
CrossRef - Chen, J., Han, G., Shang, C., Li, J., Zhang, H., Liu, F., Wang, J., Liu, H., Zhang, Y., 2015. Proteomic analyses reveal differences in cold acclimation mechanisms in freezing- tolerant and freezing-sensitive cultivars of alfalfa. Front. Plant Sci. 6, 105.
CrossRef - Fan K., Cardona C., Li Y., Shi Y., Xiang X., Shen C., et al. Rhizosphere-associated bacterial network structure and spatial distribution differ significantly from bulk soil in wheat crop fields. Soil Biol. Biochem. 2017. 113, 275–284. doi: 10.1016/j.soilbio.06020
CrossRef - Vorholt JA, Vogel C, Carlstrom CI, Muller DB. Synthetic communities in plant microbiome research. Cell Host Microbe. 2017;22:142-155
CrossRef - Matthews A., Pierce S., Hipperson H. and Raymond B. Rhizobacterial community assembly patterns vary between crop species. Front. Microbiol. 2019. 10, 581.
CrossRef - Trivedi, P., Leach, J. E., Tringe, S. G., Sa, T. & Singh, B. K. Plant-microbiome interactions: from community assembly to plant health. Nat. Rev. Microbiol. 2020.,18, 607–621.
CrossRef - Howard MM, Munoz CA, Kao-Kniffin J, Kessler A. Soil microbiomes affect crop growth. Front Plant Sci. 2020;11:1171.
CrossRef - Ling N., Wang T. and Kuzyakov Y. Rhizosphere bacteriome structure and functions. Nat Commun 13, 836. 2022. https://doi.org/10.1038/s41467-022-28448-9.
CrossRef - Beule L., Lehtsaar E., Corre M. D., Schmidt M., Veldkamp E., and Karlovsky P. Poplar rows in temperate agroforestry croplands promote bacteria, fungi, and denitrification genes in soils. Front. Microbiol. 2020. 10:3108. doi: 10.3389/fmicb.2019.03108.
CrossRef - Beule L., and Karlovsky P. Tree rows in temperate agroforestry croplands alter the composition of soil bacterial communities. PLoS One 16, 2021. e0246919–e0246920. doi: 10.1371/journal.pone.0246919
CrossRef - Ben zineb A., Barkaoui K., Karray F., Mhiri N., Sayadi S., Mliki A. and Gargouri M. Olive agroforestry shapes rhizosphere microbiome networks associated with annual crops and impacts the biomass production under low-rainfed conditions. Front. Microbiol. 13:977797. doi: 10.3389/fmicb.2022.977797
CrossRef - Beule L., Vaupel A., Moran-Rodas V.E. Abundance, Diversity, and Function of Soil Microorganisms in Temperate Alley-Cropping Agroforestry Systems: A Review. Microorganisms 2022, 10, 616. https://doi.org/10.3390/microorganisms10030616
CrossRef - Dai, CC., Chen, Y., Wang, XX. Effects of intercropping of peanut with the medicinal plant Atractylodeslancea on soil microecology and peanut yield in subtropical China. Agroforest Syst 2013., 87, 417–426. https://doi.org/10.1007/s10457-012-9563-z
CrossRef - Mondal S., Pramanik K, Pal P, Mitra S, Ghosh SK, Mondal T, Soren T, Maiti TK. Multifaceted roles of root exudates in light of plant-microbe interaction. Unravelling plant-microbe synergy. Academic press. 2023. pp 49-76.https://doi.org/10.1016/B978-0-323-99896-3.00003-5
CrossRef - Vicré M, Santaella C, Blanchet S, Gateau A, Driouich A. Root border-like cells of Arabidopsis. Microscopical characterization and role in the interaction with rhizobacteria. Plant Physiol 2005.,138:998–1008. doi:10.1104/pp.104.051813
CrossRef - De Weert S., Vermeiren H., Mulders I.H.M., Kuiper I., Hendrickx N., Bloemberg G.V., Vanderleyden J., De Mot R., Lugtenberg B.J.J. Flagella-driven chemotaxis towards exudate components is an important trait for tomato root colonization by Pseudomonas fluorescens. Mol Plant-Microb Interact 2002. 15:1173–1180. doi:10.1094/MPMI.2002.15.11.1173
CrossRef - Kwak Y., Bakker PAHM, Glandorf DCM, Topham J., Paulitz T., Weller D.M. Diversity, virulence and 2,4-diacetylphloroglucinol sensitivity of Gaeumannomyces graminis var. tritici isolates from Washington State. Phytopathol. 2009. 99:472–479. doi:10.1094/PHYTO-99-5-0472
CrossRef - Wen, T., Zhao, M., Yuan, J. et al. Root exudates mediate plant defense against foliar pathogens by recruiting beneficial microbes. Soil Ecol.2021. Lett. 3, 42–51. https://doi.org/10.1007/s42832-020-0057-z
CrossRef - Rudrappa T, Kirk J, Czymmek PW, Paré PW, Bais HP. Root-secreted malic acid recruits beneficial soil bacteria. Plant Physiol 2008. 148:1547–1556. doi:10.1104/pp.108.127613
CrossRef - Leeman M, Van Pelt JA, Hendrickx MJ, Scheffer RJ, Bakker PAHM, Schippers B . Biocontrol of fusarium wilt of radish in commercial greenhouse trials by seed treatment with Pseudomonas fluorescens WCS374. Phytopathol.1995. 85:1301–1305
CrossRef - Weller DM. Pseudomonas biocontrol agents of soilborne pathogens: looking back over 30 years. Phytopathol.2007. 97:250–256. doi:10.1094/PHYTO-97-2-0250
CrossRef - De Weger LA, Van Arendonk JJCM, Recourt K, Van der Hofstad GAJM, Weisbeek PJ, Lugtenberg B Siderophore-mediated uptake Of Fe-3+ by the plant growth-stimulating Pseudomonas putida strain WCS358 and by other rhizosphere microorganisms. J Bacteriol. 1988. 170:4693–4698
CrossRef - Fridlender M, Inbar J, Chet I. Biological control of soilborne plant-pathogens by a ß-1,3 glucanase-producing Pseudomonas cepacia. Soil Biol Biochem. 1993. 25:1211–1221
CrossRef - Ton J, Davison S, Van Wees SC, Van Loon L, Pieterse CM. The arabidopsis ISR1 locus controlling rhizobacteria-mediated induced systemic resistance is involved in ethylene signaling. Plant Physiol. 2001;125(2):652-661. doi:10.1104/pp.125.2.652
CrossRef - Bittel P, Robatzek S. Microbe-associated molecular patterns (MAMPs) probe plant immunity. Curr Opin Plant Biol. 2007;10(4):335-341. doi:10.1016/j.pbi.2007.04.021
CrossRef - Chinchilla D, Bauer Z, Regenass M, Boller T, Felix G. The Arabidopsis receptor kinase FLS2 binds flg22 and determines the specificity of flagellin perception. Plant Cell. 2006;18(2):465-476. doi:10.1105/tpc.105.036574
CrossRef

