Introduction
The rampant use of agricultural fertilizers to meet the demand of food for burgeoning population has remained a contentious issue, especially when the world is faced with serious concerns like climate change, Greenhouse effect, depleting land resources and fossil fuels. Early 1950s witnessed pioneering efforts by Norman Boralug, who introduced the Green Revolution model to address food security concerns. This model among other tenets, emphasized on the adoption of nitrogenous fertilizers in the farmers’ fields. Although the model became a pervasive aspect of agriculture, its excessive use proved unsustainable, because the application was undertaken without comprehensive and long-term assessment of their ecological, economic, and agronomic consequences.1,2 The impact of green revolution in agriculture was overwhelming as the production enhanced and mortality rate decreased, but the sustainability of Green revolution itself was at stake because enhancing agricultural production remained the main focus, while other concerns like environmental sustainability and nutritional value of the produce were overlooked.2,3 Among other environmental hazards, the nitrogen fertilizer industry and distribution chain of fertilizers contributed to 5 percent of the total nitrous oxide (a potent Greenhouse gas) emissions.4 In 2023 alone, the emissions from pre and post agricultural production of fertilizer manufacturing was estimated to be 94.26kt, 32.37 kt, 2.01 kt and 0.423 kt for China, USA, Brazil and India respectively.5 Nitrogenous fertilizers add nearly 150 million tons reactive nitrogen including HONO, NH3 and NO (nitrous acid, ammonia and nitric oxide) globally, and this is expected to increase further to 600 million tons in the coming years.6 Adding to the nitrogen footprint is also the nitrate runoff, that pollutes soil and water bodies and increases chances of blue baby syndrome or methemoglobinemia and carcinomas in children and adults respectively. The changes in the chemistry of the atmospheric air on the health of man and ecosystems costs approximately USD 340 billion to USD 3.4 trillion yearly (https://www.unep.org/news-and-stories/story/four-reasons-why-world-needs-limit-nitrogen-pollution, accessed on 29.11.2025). So, it is essential to bring down nitrogen foot print and for this, nitrogenous fertilizers must be substituted by other ecofriendly and sustainable options like biofertilizers or microbial inoculants.
Nitrogen is the most abundant gas, comprising nearly 78 percent of the total atmospheric gases, contributing to vital component of biomolecules viz. amino acids, proteins, chlorophyll, nucleic acids, nitrogenous bases, alkaloids and numerous secondary metabolites in plants. However, plants are unable to access it directly from the environment. Diazotrophs are group of nitrogen-fixing microorganisms including bacteria, archaea and blue green algae, that play crucial role in reducing atmospheric nitrogen to plant usable forms like ammonium, referred as biological nitrogen fixation (BNF) through nitrogenase enzyme, a metalloprotein complex encoded by the nifH gene.7 This process harnesses the physiological potential of nitrogen-fixers that find application in biofertilizer formulations. Diazotrophs have been isolated from the root nodules of leguminous plants, but recent years have seen their exploration from the non-legumes also. Most pioneering work on nitrogen-fixing bacteria was undertaken by Johanna Döbereiner, who originally hailed from Czechoslovakia, but later in 1953 moved to Brazil. Due to the contributions of Döbereiner and group, Brazil stands at the apex, surpassing many other countries for research on diazotrophic bacteria.8 Although, conventional nitrogen fixers like Rhizobia, Azotobacter, Azospirillum, Azolla–Anabaena symbiosis etc. routinely find application as commercial inoculants, there are a vast number of other less explored diazotrophs with multiple PGP (plant growth promoting) traits, that can be developed into bioinoculants.9 PGPR (plant growth promoting rhizobacteria) inoculants possess PGP traits like fixation of atmospheric nitrogen, solubilization of iron by siderophore production, solubilization of phosphate, synthesis of IAA etc.10 The incorporation of diazotrophic bacteria into agricultural practices is ecofriendly and sustainable, and can reduce reliance on chemical fertilizers and therefore, nitrous oxide emissions during the production and distribution chain of nitrogenous fertilizers.11
Rauvolfia tetraphylla (barachandrika or devil pepper) belongs to Dogbane family/ Apocyanaceae and serves as a validated substitute to Rauvolfia serpentina or Sarpagandha due to their comparable alkaloid profile.12, 13 The plant originated in some parts of Northern and Southern America, and it was later introduced to India, China, Nepal, Taiwan, Vietnam and some parts of Australia.14 Rauvolfia. serpentina was popularized as an antihypertensive, antimicrobial and also for the treatment of insomnia, anxiety, etc. due to the presence of reserpine, a monoterpene indole alkaloid.15 Because of its medicinal value, Rauvolfia serpentina was ruthlessly exploited by the pharmaceutical industry, which drastically brought down its numbers, resulting in its inclusion in the endangered category by the IUCN.16 Recent in silico studies have shown that leaves of R. tetraphylla L. possess antiviral properties.17 In the Bhagalpur district (25.2372° N, 86.9746° E) of Bihar, India R. tetraphylla L. is found growing abundantly in wild patches.
The seeds of Rauvolfia tetraphylla L. have poor germination inspite of 83.1% viability. Hence, multiplication through seeds and regeneration by stem cuttings is difficult due to occurrence of phenolic compounds in the seed and also limited rooting.18 Besides, reserpine extraction from roots of Rauvolfia tetraphylla requires sacrificing the whole plant, making its sustainable utilization difficult. Therefore, it is necessary to bring this plant into cultivation and application of microbial inoculants based on diazotrophs with PGP traits like IAA synthesis, iron solubilization and phosphate solubilization can lead to plant growth promotion and also increase concentration of bioactives. Mishra and Kumar have critically reviewed the interactions of rhizospheric microbes and endophytes in Rauvolfia sp. and they could be of immense potential in sustainable agriculture.19
Among the major nitrogen-fixing bacteria is Azotobacter, which is frequently used in formulations of biofertilizers, but it suffers from several pitfalls like inconsistent performance due to variations in soil physicochemical parameters and hence may not be effective in enhancing alkaloid level and ameliorating growth. Hence, exploring rhizomicrobiota of R. tetraphylla L., other than the conventional ones can provide useful insights for the development of microbial inoculant/s specially designed for this medicinal plant.
Materials and Methods
Collection of rhizospheric soil
Healthy plants of Rauvolfia tetraphylla L. growing in the Botanical Garden of University Department of Botany, Tilka Manjhi Bhagalpur University were carefully uprooted. The plants were kept in sterile polybags, with their rhizospheric soil intact and taken to the Biolab at Bihar Agricultural University, Sabour within a few hours of collection. By vigourous shaking, bulk soil was removed, and the remaining soil adhering to the root (the rhizospheric soil) was used for the isolation of diazotrophs from the rhizosphere.
Isolation of diazotrophic bacterial groups
The rhizospheric soil was obtained by vigorous shaking in 1 L 0.9 percent NaCl for 10 minutes. The contents were put on orbital shaker at 300 rpm, for 90 minutes and then centrifuged to homogenize bacterial content in the supernatant, while the soil particles pelleted at the bottom.20 Using 1 mL of the supernatant, serial dilution was carried out until reaching 10-4 dilution. Usually 10-3 or 10-4 dilutions are taken since at higher dilutions colonies are crowded and might overlap, whereas at dilutions lesser than this, the colonies may be very few to count or almost absent from a plate. Ideally, the number of colonies on the plate should be from 30-300 for statistical accuracy. Hence 10-3 or 10-4 dilutions are preferred for enumeration and isolation of bacteria.21, 22
Isolation was done on Jensen’s agar media (procured from HiMedia) supplemented with bromothymol blue (BTB).23 100µL of each dilution was spread on the petriplates containing the sterile media in the laminar flow hood. After spreading the plates were incubated in inverted position in an incubator at 28°C till the appearance of visible colonies. The incorporation of BTB as a pH indicator is similar to that in NFb (nitrogen-free bromothymol blue) media for visual screening of diazotrophs.24 Change in colour of the media around the colonies to blue colour is regarded as nitrogen-fixation.25,26 Isolates were purified and preserved in 20% glycerol stocks until further study.
Screening for PGP traits
Screening for siderophore production
For qualitative screening for siderophore production, the selected diazotrophs were spot inoculated on Chrome Azurol Sulfonate (CAS) agar media (procured from HiMedia) and incubated at 28°C for 3-5 days. Development of orange halos around bacterial colonies indicated positive siderophore production.27
Screening for IAA production
50 μL of glycerol stock was inoculated in 5ml of nutrient broth medium containing 5 mM L-tryptophan for 48 h at 28°C. Thereafter, 500 μL of freshly prepared Salkowski reagent (1 ml of 0.5 M FeCl3 in 50 ml 35% HClO4) was added to each tube and incubated in dark for 30 minutes. Development of pink colour indicated positive IAA production.28
Screening for phosphate solubilization
This was done on Pikovskaya’s agar medium (procured from HiMedia) containing tricalcium phosphate. Bacterial isolates were spot inoculated and incubated for 48 hours at a temperature of 28°C. Development of clear halo zones around isolates indicated positive phosphate solubilization.29
Morphological characterization of selected isolates
Pure cultures (48-72 h old) were studied for their colony characteristics viz. colour, shape, size, margin etc. on Jensens agar media amended with BTB.
Biochemical and molecular characterization of selected diazotrophs
Isolates showing in vitro nitrogen-fixation by their visible blue colour on Jensens agar media amended with BTB were screened for citrate utilization, lysine and ornithine utilization, urease detection, phenylalanine deamination, nitrate reduction, H2S production, and carbohydrate metabolism tests (glucose, adonitol, lactose, arabinose, and sorbitol) using HiAssorted™ Biochemical Test Kit KB002 procured from HiMedia.30
Extraction of genomic DNA from Bacterial isolates
From glycerol stocks, the bacterial isolates were revived in nutrient broth medium till the appearance of turbidity/ visible growth. The cultures were centrifuged at 10,000 rpm for 10 minutes. Following centrifugation, the supernatant was discarded, and bacterial cells were pelleted down. From the pellets, extraction of genomic DNA was performed using HiPurA® Genomic DNA Purification Kit. The protocol provided by the manufacturer was followed, and upon isolation, the quality of separated DNA sequences was checked using agarose gel electrophoresis. The genomic DNA of the isolates in the gel were visualized by UV transilluminator.
Amplification of 16S rRNA gene with 16S universal Primers
Using purified DNA as template, 16S rRNA of the isolates were amplified using primers 27F (5′-AGAGTTTGATCCTGGCTCAG-3′) and 1492R (5′-GGTTACCTTGTTACGACTT3′). The resulting amplicons were resolved on a 1.5% TAE-agarose gel. Thereafter, the amplicons were purified and sequencing was performed using ABI 3500 XL Genetic Analyzer.31 The generated sequences were allowed to overlap to get a concatenated sequence using CodonCode Aligner (Version 12.0.4). Thereafter, the assembled sequences were converted into FASTA format files and subjected to alignment in public database using the EzBioCloud server (last updated on 21.04.2025) for identification of closely related species.32 Later, comparisons were carried on NCBI BLAST server to determine the closest phylogenetic neighbour and the sequences were deposited in NCBI GenBank using the submission portal to obtain accession numbers. Alignment was done using Clustal W and phylogenetic tree was made using MEGA 12 software (version 12.0.14).33
Results and Discussion
Isolation and screening for Nitrogen Fixation
Qualitative screening for nitrogen fixation was done on BTB containing Jensens media. The production of distinct blue colour within 2-5 days of incubation by the isolates, was taken as confirmatory test for nitrogen fixation. The study yielded 79 isolates and they were designated with codes RT1 to RT79. Among them, ten isolates produced distinct blue colour on BTB containing Jensens media, but the fixation was higher among the isolates RT2, RT34, RT40A, RT71, RT77, RT78 and RT79. Fig 1. Showing high nitrogen fixation by isolates RT71, RT77 and RT78 on BTB containing Jensens media Depicted the results for in vitro screening for nitrogen fixation and PGP traits are highlighted in Table 1.
The isolate RT2 showed strong nitrogen fixation changing the colour of the Jensens medium containing BTB from forest green to blue and then to yellow and this aligns well with the earlier findings in Stenotrophomonas maltophilia.34 Stenotrophomonas maltophilia belongs to Gammaproteobacteria and earlier research has implicated their role in cystic fibrosis and also nosocomial infections, frequently leading to morbidity.35, 36, 37 In sharp contrast to these studies, several strains of S. maltophilia harbour the rhizosphere and possess PGP traits like nitrogen fixation, phosphate solubilization, production of plant growth regulators, siderophore production etc. and are also involved in biofortification and degradation of xenobiotic compounds.38 S. maltophilia have also been isolated from the roots of Arachis hypogea, where they have shown to regulate physiological and biochemical responses, and improve tolerance to stress, under limiting nitrogen conditions.39 Earlier the biofertilizer and biocontrol properties of S. maltophilia in sustainable cultivation of wheat has also been reported .40
The first report of isolation of Bacillus rugosus was done from a marine sponge.41 In another study in rice, although not reported as a diazotroph, B. rugosus occurred as an endophyte and it was found to provide tolerance to multiple stress.42 B. rugosus RT34 has shown for nitrogen fixation on BTB containing Jensens media, however further validation for nitrogen fixation would require amplification of nif gene. Among the Bacilli members isolated in this study, Bacillus stercorsis RT40A was earlier isolated from Magnaporthe oryzae in rice and tomato wilts respectively, and they were found effective in controlling blast fungus in rice and Fusarial wilts in tomato.43,44 Although not shown to be a diazotroph in previous studies, B. stercorsis RT 40A showed in vitro nitrogen fixation on BTB containing Jensens agar medium. Another PGPR, Bacillus cabrialesii was isolated from Mexican wheat varieties but its diazotrophic potential has not been reported yet.45 In sharp contrast, the diazotrophic capacity of Bacillus cabrialesii RT51 on Jensens media containing BTB has been evidenced in our study. Another member, Lysobacter prati RT71 has shown high nitrogen fixation activity and this aligns with the earlier observation in E4 Lysobacter strain.46 The earliest report of isolation of Lysobacter prati was from plateau meadow.47 The observation of nitrogen fixation by R. oryzihabitans aligns with the previous reports for the same being isolated from rice endosphere.48 Previously, Metapseudomonas lalkuanensis, was isolated from contaminated soil in Lalkuan in India .49 Although literature does not mention it as a nitrogen fixer, but our studies have shown isolate Metapseudomonas lalkuanensis RT76 to be moderately fixing-nitrogen. Nitrogen fixing ability of Pseudomonas hunanensis RT77 aligns with the nitrogen-fixing ability of the same isolate from soybean.50 Pseudomonas hunanensis SPT26 was earlier reported to secrete antifungal substances that could control wilt diseases in tomato and was proposed as promising rhizobacteria for the formulation of microbial inoculant.51 In the present study the release of siderophores by Pseudomonas hunanensis RT77 may be effective in controlling phytopathogens. Pseudomonas mosselii isolated from rice rhizosphere is a promising biocontrol agent against rice blast fungus Magnaporthe oryzae and it was also reported to enhance yield and nutrient acquisition in rice.52,53 P. mosselii isolated from durian rhizosphere was also reported as a nitrogen fixer and produced IAA from rhizosphere of durian rhizosphere.54 In our study, P. mosselii RT78 did not solubilize phosphate but showed nitrogen fixation and siderophore production in the in vitro assay. Bacillus tequilensis RT79 although showing strong nitrogen fixation in our study, has not been reported as a diatroph from previous studies.
 |
Figure 1: Petriplates showing pure cultures of RT71, RT77 and RT78 showing nitrogen fixing ability on Jensens media containing BTB |
Screening of rhizospheric diazotrophic bacteria for Plant Growth-Promoting Trait-IAA synthesis
The principal auxin, Indole-3-acetic acid (IAA) is an important phytohormone regulating plant growth and development by cell enlargement and division, tissue differentiation, and responses to light, and also controls many aspects of plant development including stimulation of root growth and nutrient acquisition from soil. The capacity to produce the phytohormone indole-3-acetic acid (IAA) is widespread among bacteria that inhabit diverse environments such as soils, fresh and marine waters, and plant and animal hosts. Among the ten diazotrophic isolates, all the isolates produced IAA, but it was significantly higher for the RT2, RT34, RT76 and RT79. This was revealed by the intensity of colour produced using Salkowski’s reagent, followed by incubation in dark. Therefore, it is expected that incorporating any of these isolates in a biofertilizer formulation will provide IAA to the host plants and improve root architecture and branching and also help in facilitating accumulation of bioactive. Enhanced IAA production by RT34 is consistent with Bacillus rugosus isolated from Arachis hypogea.55 Similarly RT2 showed higher production of IAA and this observation is in line with IAA released by S. maltophilia isolated from rhizosphere of chilli.56 Stenotrophomonas, isolated from poplar also showed auxin production in the in vitro condition similar to RT2, but there it existed as an endophyte , and in this study it was isolated from the rhizosphere.57 Higher production of IAA by Metapseudomonas lalkuanensis RT76 in this study has not been reported in any previous study. IAA production by P. mosselii RT 78 aligns with the production of IAA by the same isolate from Agave americana.58 Similarly, Bacillus tequilensis from soybean rhizosphere was shown to produce IAA , which is similar to B. tequilensis RT79 from R. tetraphylla .59 Growth responses of IAA producing rhizobacteria on dicots and monocots by pot experiments would be further required to validate their potential in agriculture.
Screening of diazotrophs for siderophore production
In soils, iron occurs mostly as Fe3+ oxy-hydroxides and must be reduced to Fe2+ to be easily assimilated by plants and microorganisms. In agriculture, this is usually accomplished by synthetic chelates, which are costly and sometimes may even lead to toxicity when used in excess.60 Besides this, alkalinity of soils in arid regions limit iron availability to plants leading to reduced crop yields, chlorosis etc. by disrupting the photosynthetic machinery. Certain rhizospheric bacteria release siderophores which help in enhancing iron availability to plants. The siderophores complexes with iron, diminishing its availability to pathogens like Fusarium oxysporum and Rhizoctonia solani, thereby exerting their plant growth promoting effect. Qualitative screening for siderophore production relied on visual examination of the halo zone around orange-yellow colonies produced by the isolates on CAS media followed by incubation for 3-5 days at 28°C. All the nitrogen fixing isolates, except RT51 and RT71 produced siderophores. S. maltophilia RT2 showed efficient iron solubilization and probably it was useful in scavenging iron from the soil.61 Rhizobium oryzihabitans RT74 also showed orange halo on CAS plate confirming siderophore production, however efficiency of solubilization is lesser as compared to other diazotrophs, and this observation is similar to earlier reports regarding siderophore production, in addition to IAA and ACC deaminase.48 Bacillus rugosus RT34 is also positive for siderophore release, and this observation is in line with the observation in rice, where the same isolate also provides tolerance to salinity and drought stress.62 Another Bacilli isolate, Bacillus tequilensis RT79 from R. tetraphylla was evidenced to solubilize iron and this observation was in line with earlier research focussing on biocontrol properties of this isolate against Fusarium and it also showed production of IAA and siderophore.63 Members of Pseudomonas generally produce siderophores under iron limiting conditions.64 M. lalkuanensis RT76 has been shown to produce siderophore P. mosselii RT78 produced siderophore and this was also observed in P. mosselii from durian rhizosphere.54 Rhizobacteria producing siderophores control phytopathogens by secreting antimicrobial compounds.65
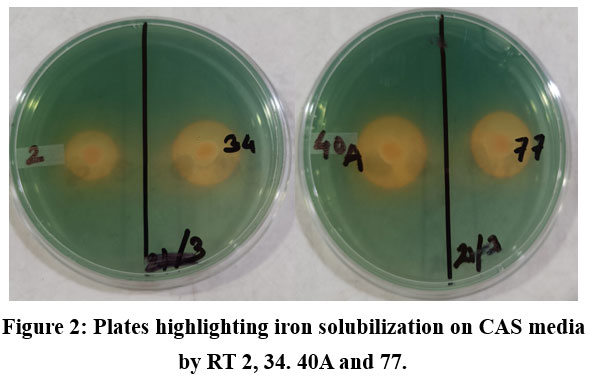 |
Figure 2: Plates highlighting iron solubilization on CAS media by RT2, RT34, RT40A and RT77. |
Screening of diazotrophs for phosphate solubilization
After nitrogen, phosphorus is the second most important macronutrient that are needed by plants for growth and development. But sometimes, phosphorus availability to plants is also limited because it occurs in combined form with metal complexes. Phosphate-solubilizing bacteria (PSB) are applied as inoculants in fields to improve bioavailability of phosphate from the soil by converting organic and insoluble forms of phosphorus into soluble forms. Some diazotrophs, in addition to fixing nitrogen, also contribute to phosphorus solubilization through the release of organic acids and enzymes, further contributing to plant growth. It has been suggested that several nitrogen-fixing strains of Azotobacter vinelandii have demonstrated promising phosphate-solubilizing capabilities and they could be used as superior candidates for biofertilizer development.66 In rice, phosphate-solubilizing diazotrophic bacteria like Herbaspirillum and Burkholderia were isolated and they were found to improve nitrogen use efficiency too.67 In our study, diazotrophic isolates RT2, RT40A, and RT51 showed solubilization zone around colonies on Pikovskaya’s agar and the efficiency of phosphate solubilization was almost comparable for isolates RT2, RT40A and RT51.
The ability of Stenotrophomonas maltophilia RT2 to solubilize phosphate aligns with previous research, where this isolate was also used for reclamation of wastes.68 S. maltophilia was earlier isolated from Porang, and it was able to fix nitrogen and also solubilize phosphate.69Among the Bacilli isolates, Bacillus stercoris RT40A and B. cabrialessi RT51 have shown phosphate solubilization but this PGP trait is not shown by the other two Bacilli isolates B. rugosus and B. tequilensis. The production of IAA and phosphate solubilization by RT40A, have previously been reported in Bacillus stercoris.44Pseudomonas hunanensis RT77 has shown no phosphate solubilization in our study, however the same isolate has shown other PGP traits in addition to phosphate solubilization in earlier studies.70 Although none of the Pseudomonas isolates have shown phosphate solubilization in our study, nevertheless phosphate solubilizing bacteria including Bacillus, Pseudomonas, Serratia and Enterobacter find application in the development of microbial inoculants and can be used to boost plant growth and development.71
 |
Figure 3: Plates showing screening for phosphate solubilization on Pikovskaya’s agar medium. Note : Although RT34 forms a big colony but the halo is missing, so it is considered negative for phosphate solubilization. |
Table 1: The table highlights PGP traits of the diazotrophs. The efficiency of each bacterial partner is indicated by + (low), ++ (moderate), or +++ (high) or ++++(very high) and the absence of PGP trait is indicated by ‘– ‘ sign.
| Bacterial Isolate | Nitrogen Fixation | Siderophore Production | Phosphate Solubilization | IAA Production |
| RT2 | ++++ | +++ | ++ | +++ |
| RT34 | +++ | +++ | – | +++ |
| RT40A | +++ | +++ | ++ | + |
| RT51 | ++ | – | ++ | ++ |
| RT71 | ++++ | – | – | + |
| RT74 | ++ | + | – | + |
| RT76 | ++ | + | – | +++ |
| RT77 | +++ | +++ | – | + |
| RT78 | +++ | ++++ | – | + |
| RT79 | +++ | ++++ | – | +++ |
Carbon utilization pattern of selected nitrogen-fixing bacterial isolates
Different isolates showed varying responses for the utilization of different carbohydrates. Except RT34, RT40A and RT74, none of the isolates were capable of utilizing glucose. Adonitol utilization was shown by isolate RT74 only. None of the isolates utilized lactose except RT74. Isolates RT34, RT40A, RT74, RT77 and RT78 showed utilization of arabinose. Only one isolate RT74 was capable of utilizing sorbitol. The results for carbon utilization pattern is highlighted in table 2.
Stenotrophomonas maltophilia RT2 was found to be non-fermentative and this is consistent with the results of Swapna and Audipudi.56 Glucose and arabinose utilization by B. rugosus is consistent with the findings in Arachis hypogea by Rabari et al.55 B. rugosus RT34 was not able to utilize lactose, but B. rugosus from Arachis hypogea was found to utilize lactose.55 One of the plausible reasons for this could be that certain bacteria prefer using simpler sugars, and may require ATP to breakdown disaccharide and therefore sticks to the former. Studies on adonitol utilization revealed that RT34 was not able to utilize adonitol, however previous studies have not revealed anything on the utilization of adonitol by this isolate. Figueroa-Brambila et al., have reported glucose fermentation by B. cabrialessi but this is not the case with B. cabrialesii RT51.72 Previous studies have not explicitly mentioned studies on sugar utilization by B. cabrialessi. Lysobacter prati RT71 did not utilize sorbitol, arabinose and lactose and these results are consistent with the previous findings.47 The ability of Bacillus stercorsis RT40A for citrate utilization and glucose fermentation is consistent while negative nitrate reduction is contrary to the findings reported so far.73 R. oryzihabitans RT74 was found to ferment arabinose, sorbitol, lactose, adonitol and glucose is in sync with the earlier findings.48 Metapseudomonas lalkuanensis RT76 did not utilize sorbitol, any sugar in our study , but it showed utilization of glucose, arabinose, lactose, adonitol, and sorbitol as earlier reports.49 Pseudomonas hunanensis RT77 and Pseudomonas mosselli RT78 utilized only arabinose and not any other sugars and this is a characteristic for several Pseudomonads . Bacillus tequilensis RT79 did not utilize any of the carbon sources tested including glucose, sorbitol, adonitol, lactose and arabinose and this observation is in sharp contrast to earlier studies indicating that phenotypic variability could be common.74
Table 2: Shows the ability of diazotrophs to utilize various carbon sources.
| Glucose | Adonitol | Lactose | Arabinose | Sorbitol | |
| RT 2 | – | – | – | – | – |
| RT 34 | + | – | – | + | – |
| RT 40A | + | – | – | + | – |
| RT 51 | – | – | – | – | – |
| RT 71 | – | – | – | – | – |
| RT 74 | + | + | + | + | + |
| RT 76 | – | – | – | – | – |
| RT 77 | – | – | – | + | – |
| RT 78 | – | – | – | + | – |
| RT 79 | – | – | – | – | – |
Biochemical Characterization of the selected diazotrophs
All the isolates were capable of utilizing lysine as the sole nitrogen source except RT71. Citrate utilization was shown by the isolates RT34, RT40A, RT71, RT74, RT76, RT77, RT78 and RT79. All the isolates utilized ornithine except RT71 and RT79. All the isolates showed urease activity except RT71. None of the isolates were capable of phenylalanine deamination. Nitrate reduction was shown only by the isolates RT74 and RT78. Only RT77 produced H2S. Overall, the biochemical profiling of the isolates highlight their functional potential in the rhizosphere and are in agreement with the earlier findings, and minor variations have also been observed. In Table 3 biochemical characterization of the selected nitrogen fixers are mentioned.
S. maltophilia was found positive for nitrate reduction, but in our study it did not show nitrate reduction.75 Further, urease positive result by RT2 is found to be consistent with earlier studies. Similarly, positive responses to lysine, ornithine, urease activity and negative for phenylalanine deaminase utilization is consistent with the results.56 While negative nitrate reduction in RT2 is in sharp contrast to positive tests for nitrate reduction by Stenotrophomonas, probably indicating that nitrate reduction may not be a useful feature of all Stenotrophomonas isolates.56 Citrate utilization by Bacillus stercorsis RT40A is consistent with earlier findings while negative nitrate reduction is contrary to the earlier findings reported.73
B. cabrialesii RT51 in sharp contrast to previous studies was neither able to reduce nitrite to nitrate, nor able to use citrate, which possibly may be due to strain variations.45 Negative lysine utilization, ornithine utilization and urease activity is consistent, while negative nitrate reduction of L. prati RT71 is not in sync with the observation of Fang et al.47 R. oryzihabitans RT74 showing ornithine utilization, urease activity, nitrate reduction is similar to the observation.48 Metapseudomonas lalkuanensis RT76 utilized citrate as earlier but did not show nitrate reduction in sharp contrast to earlier studies by Thorat et al. where it was positive for nitrate reduction.49 P. hunanensis RT77 showed positive citrate utilization, negative nitrate reduction and this was consistent with the reports of Gao et al.76 P. mosselii RT78 showed similar traits to P. hunanensis except negative nitrate reduction and positive hydrogen sulfide production for the former, and vice versa for former. Bacillus tequilensis RT79 utilized citrate, ornithine, lysine, showed urease activity and nitrate reduction, and this is similar to the observation of Gatson et al.74
Table 3: Shows biochemical characterization of the selected nitrogen fixers
| Citrate utilization | Lysine Utilization | Ornithine utilization | Urease Activity | Phenylalanine Deamination | Nitrate Reduction | H2S Production | |
| RT2 | – | + | + | + | – | – | – |
| RT34 | + | + | + | + | – | – | – |
| RT40A | + | + | + | + | – | – | – |
| RT51 | – | + | + | + | – | – | – |
| RT71 | + | – | – | – | – | – | – |
| RT74 | + | + | + | + | – | + | – |
| RT76 | + | + | + | – | – | – | – |
| RT77 | + | + | + | + | – | – | + |
| RT78 | + | + | + | + | – | + | – |
| RT79 | + | + | – | + | – | – | – |
Morphological traits of selected isolates
For all the isolates, colonies were round with smooth consistency, shape and margins. Certain isolates exceeded 1mm in diameter, and for some they were less than 1mm. Colour of colonies were generally white except RT34 and RT74, which were yellow in colour. The isolates were all rod shaped, and Gram negative, except Bacilli isolates RT34, RT40A, RT51 and RT79, which were Gram positive. The colonies of the diazotrophs were mostly slimy or mucoid, probably a mechanism to prevent nitrogenase enzyme from inactivation due to oxygen. Colony morphology and Grams reaction were similar to previous findings for all the isolates. The morphological characters of selected diazotrophs are mentioned in Table 4.
Table 4: Shows morphological characteristics of the selected diazotrophs.
| Colony shape | Colony size | Margin of colony | Colony elevation | Chromogenesis | Surface of Colony | Texture | Cell Shape | Grams Staining | |
| RT2 | Round | >1mm | Entire | Raised | White to pale yellow | Smooth | Mucoid | Rod | Negative |
| RT34 | Round | >1mm | Entire | Convex | yellow | Rugose | Butyrous | Rod | Positive |
| RT40A | Round | >1mm | Undulate | Convex | Creamy white | Wrinkled | Rough and dry | Rod | Positive |
| RT51 | Round | >1mm | Entire | Convex | White | Rough | Slimy | Rod | Positive |
| RT71 | Round | <1mm | Entire | Raised | Off white | Smooth and shiny | Mucoid | Rod | Negative |
| RT74 | Round | >1mm | Entire | Raised | Yellow | Smooth and shiny | Slimy | Rod | Negative |
| RT76 | Round | >1mm | Entire | Flat | Round | Smooth and moist | Mucoid | Rod | Negative |
| RT77 | Round | >1mm | Entire | Flat | White | Smooth and moist | Slimy | Rod | Negative |
| RT78 | Round | >1mm | Entire | Flat | White | Smooth and moist | Slimy | Rod | Negative |
| RT79 | Round | >1mm | Entire | Convex | White | Rough | Mucoid | Rod | Positive |
Molecular Phylogenetic Identification of Diazotrophs by 16S rRNA gene sequencing
Molecular phylogenetic diversity studies of diazotrophs by 16S r RNA gene sequencing, confirmed four species of Bacilli, three Pseudomonads, one each of Lysobacter, Stenotrophomonas and Rhizobium, showing 100% sequence similarity with the corresponding strains on NCBI. Fig 4: showed amplification of conserved 16S rRNA gene of the rhizospheric isolates, with M indicating 1 kb marker. Table 5 below highlights 16S rRNA based identification of diazotrophs using EzBiocloud and NCBI. NCBI Accession numbers of the isolates are also mentioned. Fig 5 – In fig 5 phylogenetic tree showing isolated diazotrophic strains from the rhizosphere of R. tetraphylla L. have been depicted. Bootstrap values are indicated on the branches and scale bar showing the number of substitutions per site is indicated below the tree.
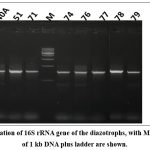 |
Figure 4: Amplification of 16S rRNA gene of the diazotrophs, with M indicating marker of 1 kb DNA plus ladder are shown. |
Table 5: Molecular identification of diazotrophs based on EzBiocloud type strains and NCBI reference sequences.
| Isolate code | Homologous microorganism(% identity)(Accession no. of NCBI reference seq.) | Identified organism | GenBank Accession Number |
| RT2 | Stenotrophomonas maltophilia M 15 (100%) ( LT222224.1) |
Stenotrophomonas maltophilia RT2 | PX363496.1 |
| RT34 | Bacillus rugosus strain SPB7(100%)(NR_181236.1) | Bacillus rugosus RT34 | PX363497.1 |
| RT40A | Bacillus stercoris strain JCM 30051 (100%) (MN536904.1) | Bacillus stercoris RT40A | PX363498.1 |
| RT51 | Bacillus cabrialesii strain TE3 (100%) ( MK462260.1 ) | Bacillus cabrialessi RT51 | PX363499.1 |
| RT71 | Lysobacter prati SYSU H10001 (Type) (100%)(MN181427) | Lysobacter prati RT71 | PX363500.1 |
| RT74 | Rhizobium oryzihabitans strain M15 (100%) MT023790.1 | Rhizobium oryzihabitans RT74 | PX363501.1 |
| RT76 | Metapseudomonas lalkuanensis strain PE08(100%) (MF943158.1 ) | Metapseudomonas lalkuanensis RT76 | PX363502.1 |
| RT77 | Pseudomonas hunanensis strain LV(100%)( JX545210.1) | Pseudomonas hunanensis RT77 | PX363503.1 |
| RT78 | Pseudomonas mosselii strain CIP 105259 (100%) (AF072688.2) | Pseudomonas mosselii RT78 | PX363504.1 |
| RT79 | Bacillus tequilensis KCTC13622 (100%)( AYTO01000043.1) | Bacillus tequilensis RT79 | PX363505.1 |
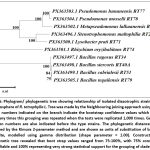 |
Figure 5: Phylogram/ phylogenetic tree showing relationship of isolated diazotrophic strains from the rhizosphere of R. tetraphylla L. |
Tree was made by the Neighbouring joining approach using MEGA 12. The numbers indicated on the branch indicate the bootstrap confidence values which means how many times this grouping was repeated when the tests were replicated 1,000 times. GenBank accession numbers are also indicated before the type strains. The phylogenetic distances were calculated by the Kimura 2-parameter method and are shown as units of substitution of bases at each site, modelled using gamma distribution (shape parameter = 1.00). Construction of phylogenetic tree revealed that bootstrap values ranged from 75-100%, with 75% considered reliable and 100% representing very strong statistical support for the grouping of clades.
The earliest derived lineage is represented by the Bacilli clade, while the other taxa were derived later. Basal taxon is Bacillus tequilensis RT79 meaning it diverged earliest as compared to other sister taxa. The diazotrophs based on Bacilli (Phylum – Firmicutes) clustered together in the same clade and while the Pseudomonads (Phylum – Gammaproteobacteria) clustered in a separate clade. Two other diazotrophs, Stenotrophomonas maltophilia RT2 and Lysobacter prati RT71, belonging to Gammaproteobacteria have also been isolated and they form a separate clade showing 100% bootstrap value. Two Pseudomonas isolates RT77 and RT78 clustered together in the same clade showing 100% bootstrap values. Metapseudomonas lalkuanensis RT76 and the two isolates Pseudomonas hunanensis RT77 and P. mosselli RT78 were derived from a common ancestor, again showing consistent 100% bootstrap value. Rhizobium oryzihabitans RT74 (Alphaproteobacteria) and all the members of Gammaproteobacteria/ Pseudomonads evolved from a common ancestor and this is supported by a strong bootstrap value of 100%.
Conclusion
The rhizosphere of R. tetraphylla L. harboured immense diversity of diazotrophs showing multiple PGP traits. A total of 79 rhizospheric isolates were obtained on Jensen’s agar media and among these 10 isolates showed positive nitrogen-fixing capacity indicated by their ability to change forest green colour to blue on BTB containing Jensens media. The rhizospheric isolates from R. tetraphylla L. belonged to Bacilli, Pseudomonads, Lysobacter, Stenotrophomonas and Rhizobium. A careful analysis of Table 1 indicates that RT2 and RT40A are showing all the PGP traits including nitrogen fixation, siderophore production, phosphate solubilization and IAA synthesis. Although RT2 and RT40A showed comparable PGP traits, but IAA production by RT2 is comparatively higher than RT40A. Hence, based on PGP traits highlighted in table 1, we recommend that Stenotrophomonas maltophilia RT2 is the best isolate and it can be further studied for its colonization and plant growth promoting effects in in vivo conditions.
Stenotrophomonas maltophilia RT2 strain is expected to provide increased levels of fixed nitrogen to Rauvolfia plants that can be assimilated into biomolecules including monoterpene indole alkaloids(reserpine), nitrogenous bases, amino acids, proteins and nucleic acids, enhancing root and shoot biomass and also accommodating elevated concentration of alkaloids. Iron is an important micronutrient in plants, and also a component of electron transport system (cytochromes), chlorophyll and ferredoxin etc., and plants need to maintain iron homeostasis. Enhanced IAA release by RT2 in the rhizosphere of R. tetraphylla is further expected to increase root branching and its architecture, to provide home for the biosynthesis and accumulation of alkaloids. Hence, developing an microbial inoculant based on RT2 is expected to manifest overall health benefits to Rauvolfia. Screening for phosphate solubilization on Pikovskaya’s agar medium has explicitly revealed the potential of RT2 for phosphate solubilization, and therefore developing inoculants based on RT2 can help in improving phosphate flux into molecules like ATP, NADPH, phospholipids, nucleic acid backbone etc. Overall such studies can be useful in addressing sustainable cultivation of not only medicinal plants but also other crops. Whole genome sequencing studies of RT 2 should also be performed to know about the presence of gene/s and gene products responsible for PGP activities. Besides, since the isolate has previously been reported to cause nosocomial infections, it would be necessary to carry out functional genomic studies to ascertain its pathogenicity in humans. Besides, annotation of genes in whole genome of the organism would be useful in identifying other potential uses of this bacteria like production of secondary metabolites. Further studies are required to test the efficiency of this isolate by the acetylene reduction assay. Various combinations of isolates can be made into microbial consortia to study microbial synergies in plant growth promotion. Besides this, assessment of increase in alkaloid level following inoculation into tissue culture raised R. tetraphylla L. plants to ascertain increment in the alkaloid level could be beneficial to the pharmaceutical industry, which at present faces scarcity of raw materials due to their over exploitation and switching to substitutes. Thus, isolation and identification of diazotrophs particularly unconventional ones, with excellent PGP traits, from the rhizosphere of R. tetraphylla points out that they could be complementing the conventional nitrogen-fixing microbial partners by adapting to diverse ecological niches. These endeavours would be valuable in addressing SDG 2030 by reducing excessive reliance on chemical fertilizers, and promote ecological sustainability in the production of medicinal plants and bioactives.
Acknowledgement
The present work was conducted at the Biolab, Bihar Agricultural University (BAU), Sabour, Bhagalpur. We sincerely thank Prof. Anshuman Kohli, Director, Department of Soil Science and Agricultural Chemistry, for granting permission to carry out this research. Our gratitude extends to Dr. Rajiv Rakshit, Associate Professor and Senior Scientist at Bihar Agricultural University, Sabour for providing necessary laboratory facilities that were instrumental in the successful completion of this work.
The authors also sincerely wish to express their heartfelt thanks to Biologia Research Pvt. Ltd., Karnal, Haryana for their invaluable co-operation in the smooth completion of all sequencing-related work.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The authors do not have any conflict of interest.
Data Availability Statement
16S rRNA gene sequences of the various diazotrophs have been deposited in the GenBank database and can be retrieved from their accession numbers.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Permission to reproduce material from other sources
Not Applicable
Authors’ contribution
Manisha Mishra: Study design, conceptualization, methodology, data analysis, investigation, writing-original draft, revision and editing
Sandeep Kumar: Study design, methodology, data analysis, revision and editing
Arvind Kumar: Study design, methodology and supervision
References
- Eliazer Nelson ARL, Ravichandran K, Antony, U. The impact of the Green Revolution on indigenous crops of India. Ethn. Foods. 2019; 6(1): 1-10.https://doi.org/10.1186/s42779-019-0011-9
CrossRef - John DA and Babu Lessons from the aftermaths of green revolution on food system and health. Front. Sustain. Food Syst. 2021;5:1-6 https://doi.org/10.3389/fsufs.2021.644559
CrossRef - Davis KF, Chhatre A, Rao ND, Singh D, Ghosh-Jerath S, Mridul A, … & DeFries R. Assessing the sustainability of post-Green Revolution cereals in India. Natl. Acad. Sci. U.S.A. 2019; 116(50) : 25034-25041. https://doi.org/10.1073/pnas.1910935116
CrossRef - Gao Y, Cabrera Serrenho A. Greenhouse gas emissions from nitrogen fertilizers could be reduced by up to one-fifth of current levels by 2050 with combined interventions. Nat. Food. 2023; 4(2) :170-178. https://doi.org/10.1038/s43016-023-00698-w
CrossRef - Food and Agriculture Organization of the United Nations. Soil Nutrient Budget. Available online: https://www.fao.org/faostat/en/#data/ESB (accessed on 05.11.2025)
- Ren C, Huang X, Wang Y, Zhang L, Zhou X, Sun W … Wang T. Enhanced Soil Emissions of Reactive Nitrogen Gases by Fertilization and Their Impacts on Secondary Air Pollution in Eastern China. Sci. Technol. 2025; 59(10) : 5119-5130. https://doi.org/10.1021/acs.est.4c12324
CrossRef - Si Z, Wang C, Zhao M, Ji Z, Qiao Y, Wang L. Genome-Wide Characterization of Nitrogenase Reductase (nifH) Genes in the Sweet Potato [Ipomoea batatas (L.) Lam] and Its Wild Ancestors. Genes (Basel). 2022; 13(8):1428. https://doi.org/10.3390/genes13081428
CrossRef - Thiebaut F, Urquiaga MCDO, Rosman AC, da Silva ML, Hemerly AS. The impact of non-nodulating diazotrophic bacteria in agriculture: understanding the molecular mechanisms that benefit crops. J. Mol. Sci. 2022; 23(19) : 11301. https://doi.org/10.3390/ijms231911301
CrossRef - Kumar U, Kaviraj M, Rout S, Singha A, Padbhushan R. Nutrient and pest-managing microbial inoculants for sustainable production of rice. In : Arora NK, Bouizgarne B, Microbial Biotechnology for Sustainable Agriculture. Springer Nature Singapore; 2024 : 331-367. https://doi.org/10.1007/978-981-97-2355-3_13
CrossRef - Kloepper Plant growth-promoting rhizobacteria on radishes. In : Proc. of the 4th Internet. Conf. on Plant Pathogenic Bacter, Station de Pathologie Vegetale et Phytobacteriologie, INRA, Angers, France. 1978 ; Vol. 2 : 879-882 (Accessed on 25.12.2025)
- https://resoilfoundation.org/en/circular-bioeconomy/fao-efficient-nitrogen-report/, (accessed on 31.10.2025)
- Shenya S, Dinara S, Sohani M. Phytochemical composition, medicinal applications, and pharmacological insights: a comparative review of Rauvolfia serpentina and Rauvolfia tetraphylla. Asian Plant Research Journal. 2025; 13(1) : 46-55. https://doi.org/10.9734/aprj/2025/v13i1292
CrossRef - Sulaiman CT, Jyothi CK, Unnithan JK, Prabhukumar KM, and Balachandran I. Identification of suitable substitute for Sarpagandha (Rauvolfia serpentina (L.) Benth. ex-Kurz) by phytochemical and pharmacological evaluation. 2020; , 9 : 1-11. https://doi.org/10.1186/s43088-020-00069-5
CrossRef - POWO, 2025 : Plants of the World Online. Facilitated by the Royal Botanic Gardens, Kew. https://powo.science.kew.org/, (accessed on 01.11.2025).
- Vakil A clinical trial of Rauvolfia serpentina in essential hypertension. Br. Heart J. 1949; 11(4):350-5. doi: 10.1136/hrt.11.4.350.
CrossRef - Rohela GK, Bylla P, Pendli S, Korra R, Gandu R, Reuben C. High performance liquid chromatography based quantification of reserpine in Rauvolfia tetraphylla L. and enhanced production through precursor feeding. Acta Chromatogr. 2021; 34(2):120-129. https://doi.org/10.1556/1326.2021.00888
CrossRef - Ramakrishnan G, Gujjula KR, Mekala JR, Sai ST, Thanay ASR, Praveen, T, Priyanka,… Varakala NR. Exploring bioactive compounds of Rauvolfia tetraphylla(RT) for 3CLprotease of SARS-CoV2: GC-MS analysis and in-silico studies. Cell Biochem Biophys. 2024; 82 : 3383-3393. https://doi.org/10.1007/s12013-024-01421-7
CrossRef - Hoque R, Borna RS, Hoque MI, Sarker RH. In vitro plant regeneration of Rauvolfia tetraphylla: a threatened medicinal plant. PTC&B. 2020; 30(1) : 33-45. https://doi.org/10.3329/ptcb.v30i1.47789
CrossRef - Mishra M, Kumar A. Harnessing the Potential of beneficial microbes of Rauvolfia species in sustainable agricultural practices: A Review. 2025; https://doi.org/10.61289/jibs2025.02.09.0330
- Barillot CD, Sarde CO, Bert V, Tarnaud E, Cochet N . A standardized method for the sampling of rhizosphere and rhizoplan soil bacteria associated to a herbaceous root system. Microbiol. 2013; 63(2) : 471-476. https://doi.org/10.1007/s13213-012-0491-y
CrossRef - O’Toole GA. Classic spotlight: plate counting you can count on. Bacteriol. 2016; 198 (23) : 3127-3127. https://doi.org/10.1128/jb.00711-16
CrossRef - Zuberer DA. Recovery and enumeration of viable bacteria. In : Methods of Soil Analysis: Part 2. Microbiological and Biochemical Properties (eds: Weaver RW, Angle S, Bottomley P, Bezdice D, Smith S, Tabatabai A, Wollum 1994.; 5 : 119-144. https://doi.org/10.2136/sssabookser5.2.c8
CrossRef - Jain D, Sharma J, Kaur G, Bhojiya AA, Chauhan S, Sharma V, … Maharjan E. Phenetic and molecular diversity of nitrogen fixating plant growth promoting Azotobacter isolated from semiarid regions of India. Biomed Res. Int. 2021 ; 1: 6686283. https://doi.org/10.1155/2021/6686283
CrossRef - Baldani JI, Reis VM, Videira SS, Boddey LH, Baldani VLD. The art of isolating nitrogen-fixing bacteria from non-leguminous plants using N-free semi-solid media: a practical guide for microbiologists. Plant soil. 2014; 384(1):413-431. https://doi.org/10.1007/s11104-014-2186-6
CrossRef - Yadav A, Boruah JLH, Geed SR, Sharma RK, Saikia R. Occurrence, identification and characterization of diazotrophic bacteria from aerial roots of Rhynchostylis retusa (L.) Blume for plant growth-promoting activity. Arch Microbiol. 2023; 205: 131. https://doi.org/10.1007/s00203-023-03458-3
CrossRef - Döbereiner J. Isolation and identification of root associated diazotrophs. Plant soil. 1988; 110(2) : 207-212. https://doi.org/10.1007/BF02226800
CrossRef - Schwyn B, Neilands J. Universal chemical assay for the detection and determination of siderophores. Biochem. 1987; 160:47–56. https://doi.org/10.1016/0003-2697(87)90612-9
CrossRef - Bric JM, Bostock RM, Silverstone SE. Rapid in situ assay for indole acetic acid production by bacteria immobilized on nitrocellulose membrane. Appl Environ Microbiol. 1991; 57: 535–538. https://doi.org/10.1128/aem.57.2.535-538.1991
CrossRef - Pikovskaya R. Mobilization of phosphorus in soil in connection with vital activity of some microbial species. Mikrobiologiya. 1948; 17: 362–370. (accessed on 20.01.2026)
- Barnwal SK, Saleh AM. Antibiotic resistance patterns of environmental bacteria from sewage water in Vellore, India: isolation, virulence analysis, and characterization. Frontiers in Microbiology. 2025; 16 : 1640369.
CrossRef - Kumar A, Singh R, Yadav A, Giri DD, Singh PK, Pandey KD. Isolation and characterization of bacterial endophytes of Curcuma longa 3 Biotech. 2016; 6(1) : 60. https://doi.org/10.1007/s13205-016-0393-y
CrossRef - Chalita M, Kim YO, Park S, Oh HS, Cho JH, Moon J … Chun J. EzBioCloud: a genome-driven database and platform for microbiome identification and discovery. Int J Syst Evol Microbiol. 2024; 74(6). https://doi.org/10.1099/ijsem.0.006421
CrossRef - Kumar S, Stecher G, Suleski M, Sanderford M, Sharma S, Tamura K. Molecular Evolutionary Genetic Analysis version 12 for adaptive and green computing. Biol. Evol. 2024; 41: 1-9 . https://doi.org/10.1093/molbev/msae263
CrossRef - Amal AA, Osman NH. Isolation, Identification And Evaluation of The Plant Growth Promoting Activities of Endophytic Stenotrophomonas maltophilia to Stimulate Growth of Clover Plants Under Salt Stress. Res. 2022; 32(8):7-20. https://journalmrji.com/index.php/MRJI/article/view/1336. Accessed on 20.12.2025.
CrossRef - Terlizzi V, Tomaselli M, Giacomini G, et al. Stenotrophomonas maltophilia in people with Cystic Fibrosis: a systematic review of prevalence, risk factors and management. Eur J Clin Microbiol Infect Dis. 2023; 42(11):1285-1296. https://doi: 10.1007/s10096-023-04648-z.
CrossRef - Brooke JS. Stenotrophomonas maltophilia: an emerging global opportunistic pathogen. Clinical microbiology reviews. 2012; 25(1), 2-41. https://doi.org/10.1128/cmr.00019-11
CrossRef - Insuwanno W, Kiratisin P, Jitmuang A. Stenotrophomonas maltophiliaInfections: Clinical Characteristics and Factors Associated with Mortality of Hospitalized Patients. Infect Drug Resist. 2020; 13:1559-1566. https://doi: 10.2147/IDR.S253949.
CrossRef - Ghosh R, Chatterjee S and Mandal NC. In : Stenotrophomonas. In Amaresan N, Senthil Kumar M., … Sankaranarayanan A. eds. Beneficial microbes in agro-ecology.2020; 427-442. Academic Press. https://doi.org/10.1016/B978-0-12-823414-3.00020-4
CrossRef - Alexander A, Singh VK, Mishra A, et al. Plant growth promoting rhizobacterium Stenotrophomonas maltophilia BJ01 augments endurance against N2 starvation by modulating physiology and biochemical activities of Arachis hypogea. Plos one. 2019; 14(9) : e0222405. https://doi.org/10.1371/journal.pone.0222405
CrossRef - Sharma P, Pandey R and Chauhan NS. Biofertilizer and biocontrol properties of Stenotrophomonas maltophilia BCM emphasize its potential application for sustainable agriculture. Frontiers in Plant Science. 2024; 15: p.1364807. https://doi.org/10.3389/fpls.2024.1364807
CrossRef - Bhattacharya D, Villalobos de los S, Ruiz VV, et al.. Bacillus rugosus nov. producer of a diketopiperazine antimicrobial, isolated from marine sponge Spongia officinalis L. Antonie Van Leeuwenhoek. 2020; 113(11) : 1675-1687. https://doi.org/10.1007/s10482-020-01472-9
CrossRef - Thampi M, Anjali C, Vijayan S and Jisha MS. Unveiling Bacillus rugosus CRI: A multi-stress tolerant endophyte revolutionizing rice resilience. Physiological and Molecular Plant Pathology. 2024; 134: 102462.https://doi.org/10.1016/j.pmpp.2024.102462
CrossRef - Xu Q, Shan Z, Yang Z, et al. Biocontrol Potential of Bacillus stercoris Strain DXQ-1 Against Rice Blast Fungus Guy11. Microorganisms. 2025; 13(7) : 1538.https://doi.org/10.3390/microorganisms13071538
CrossRef - Pengproh R, Thanyasiriwat T, Sangdee K, et al. Evaluation and genome mining of Bacillus stercoris isolate B. PNR1 as potential agent for Fusarium wilt control and growth promotion of tomato. The Plant Pathology Journal 2023; 39(5): 430. doi: 5423/PPJ.OA.01.2023.0018
CrossRef - Figueroa-Brambila KM, Escalante-Beltrán A, Montoya-Martínez AC, et al. Bacillus cabrialesii: five years of research on a novel species of biological control and plant growth-promoting bacteria. Plants, 2023; 12 (13) : 2419. https://doi.org/10.3390/plants12132419
CrossRef - Iwata K, Azlan A, Yamakawa H & Omori T. Ammonia accumulation in culture broth by the novel nitrogen-fixing bacterium, Lysobacter E4. Journal of bioscience and bioengineering. 2010; 110 (4) : 415-418.https://doi.org/10.1016/j.jbiosc.2010.05.006
CrossRef - Fang BZ, Xie YG, Zhou XK et al. Lysobacter prati nov., isolated from a plateau meadow sample. Antonie van Leeuwenhoek. 2020; 113:763–772 https://doi.org/10.1007/s10482-020-01386-6
CrossRef - Zhao J, Zhao X, Wang J et al. Isolation, identification and characterization of endophytic bacterium Rhizobium oryzihabitans nov., from rice root with biotechnological potential in agriculture. Microorganisms. 2020; 8(4) : 608. https://doi.org/10.3390/microorganisms8040608
CrossRef - Thorat V, Kirdat K, Tiwarekar B, et al. Pseudomonas lalkuanensis nov., isolated from a bacterial consortia of contaminated soil enriched for the remediation of e-waste. International Journal of Systematic and Evolutionary Microbiology. 2020: 70 (12), : 6468-6475. https://doi.org/10.1099/ijsem.0.004559
CrossRef - Wang DD, Sun L and Yu H. Isolation of p-hydroxybenzoic acid degrading bacteria from soybean rhizosphere and their growth promoting functions. 2023; 51(20) : 11-14. 3969/j.issn.0517-6611.2023.20.003
- Verma P, Bhattacharya A, Bharti C et al.Antifungal metabolites produced by Pseudomonas hunanensis SPT26 effective in biocontrol of fusarium wilt of Lycopersicum esculentum under saline conditions. World J Microbiol Biotechnol . 2024; 40: 305 https://doi.org/10.1007/s11274-024-04092-x
CrossRef - Wu L, Xiao W, Chen G. et al. Identification of Pseudomonas mosselii BS011 gene clusters required for suppression of rice blast fungus Magnaporthe oryzae. Journal of biotechnology. 2018; 282:1-9. https://doi.org/10.1016/j.jbiotec.2018.04.016
CrossRef - Sultana R, Jashim AII, Islam SMN, et al. Bacterial endophyte Pseudomonas mosselii PR5 improves growth, nutrient accumulation, and yield of rice (Oryza sativa) through various application methods. BMC Plant Biol. 2024; 24 :1030 https://doi.org/10.1186/s12870-024-05649-6
CrossRef - Vân Trần T, Võ LT, Nguyễn LTN, et al. Identification and characterization of Pseudomonas isolated from durian growing soil in the Central Highlands, Vietnam. Tạp chí Nông nghiệp và Phát triển. 2025; 24(3) : 78-94. https://doi.org/10.52997/jad.8.03.2025
CrossRef - Rabari A, Ruparelia J, Jha CK, et al. Evaluating the Phytohormone Proficiencies of Multifarious Bacillus rugosus for Growth Promotion in Arachis hypogaea (L.). Journal of Basic Microbiology. 2025; p.e70011. https://doi.org/10.1002/jobm.70011
CrossRef - Kumar NP, Audipudi AV. Exploration of a novel plant growth promoting bacteria Stenotrophomonas maltophilia AVP27 isolated from the chilli rhizosphere soil. International Journal of Engineering Research and General Science. 2015; 3(1): 265-273. https://pnrsolution.org/Datacenter/ Vol3/Issue1/34.pdf (Accessed on 20.12.2025).
- Ulrich K, Kube M, Becker R, Schneck V, Ulrich, A. Genomic analysis of the endophytic Stenotrophomonas strain 169 reveals features related to plant-growth promotion and stress tolerance. Frontiers in microbiology. 2021; 12: p.687463.
CrossRef - Torre-Ruiz DL, Ruiz-Valdiviezo N, Rincón-Molina VM, Rodríguez-Mendiola CI et al. Effect of plant growth-promoting bacteria on the growth and fructan production of Agave americana Braz. J. Microbiol.. 2016; 47 : 587-596. https://doi.org/10.1016/j.bjm.2016.04.010
CrossRef - Kang SM, Khan AL, Waqas M, et al. Integrated phytohormone production by the plant growth-promoting rhizobacterium Bacillus tequilensisSSB07 induced thermotolerance in soybean. Plant Interact. 2019 ; 14(1) :416–423. https://doi.org/10.1080/17429145.2019.1640294
CrossRef - Sareh R, Majid T, Bahram B, et al. The role of plant growth-promoting rhizobacteria (PGPR) in improving iron acquisition by altering physiological and molecular responses in quince seedlings. Plant Physiol. Biochem. 2020; 155 : 406-415. https://doi.org/10.1016/j.plaphy.2020.07.045
CrossRef - Brooke JS. Advances in the Microbiology of Stenotrophomonas maltophilia. Clin Microbiol Rev. 2021; 34:10.1128/cmr.00030-19. https://doi.org/10.1128/cmr.00030-19
CrossRef - Thampi M, Anjali C, Vijayan S, et al. Unveiling Bacillus rugosus CRI: A multi-stress tolerant endophyte revolutionizing rice resilience. Physiological and Molecular Plant Pathology. 2024; 134 : 102462. https://doi.org/10.1016/j.pmpp.2024.102462
CrossRef - Kang SM, Khan AL, Waqas M, et al. Integrated phytohormone production by the plant growth-promoting rhizobacterium Bacillus tequilensisSSB07 induced thermotolerance in soybean. Journal of Plant Interactions. 2019 ; 14(1) :416–423. https://doi.org/10.1080/17429145.2019.1640294
CrossRef - Cézard C, Farvacques N, Sonnet P. Chemistry and biology of pyoverdines, Pseudomonas primary siderophores. Current medicinal chemistry. 2015; 22(2), 165-186.(accessed on 20.01.2026).
CrossRef - Ji SH, Gururani MA, Chun SC. Isolation and characterization of plant growth promoting endophytic diazotrophic bacteria from Korean rice cultivars. Res. 2014; 169(1) : 83-98. https://doi.org/10.1016/j.micres.2013.06.003
CrossRef - Nosrati R, Owlia P, Saderi H, et al. Phosphate solubilization characteristics of efficient nitrogen-fixing soil AzotobacterIran J Microbiol. 2014 ; 6 (4) : 285. https://pmc.ncbi.nlm.nih.gov/articles/ PMC4367947/ (Accessed on 20.12.2025).
- Estrada GA, Baldani VLD, de Oliveira DM, et al., Selection of phosphate-solubilizing diazotrophic Herbaspirillum and Burkholderia strains and their effect on rice crop yield and nutrient uptake. Plant soil. 2013; 369 (1) : 115-129. https://doi.org/10.1007/s11104-012-1550-7
CrossRef - Zhu X, Gong W, Li W, et al., Reclamation of waste coal gangue activated by Stenotrophomonas maltophilia for mine soil improvement: Solubilizing behaviour of bacteria on nutrient elements. Environ. Manage. 2022; 320: p.115865. https://doi.org/10.1016/j.jenvman.2022.115865
CrossRef - Masniawati A, Johannes E, Yusran Z, et al., Sustainable Porang Farming through Exploration of Cellulolytic-Hydrolysing, Nitrogen-Fixing, and Phosphate-Solubilizing Bacteria: Physiological and Biomolecular Analysis. Bull. 2025; 52(1) : p.29. https://doi.org/10.1134/S1062359024611741
CrossRef - Ullah A, Ahmad S, Azeem MA, Ullah S, Ullah S, Laiq M. … & Shao H Microbial Dream Team: Cotton Performs Better in Salty Soil with Pseudomonas hunanensis RT-12 and Staphylococcus caprae DS-2. Microbiol. 2026 ; 83(1) : 31. https://doi.org/10.1007/s00284-025-04614-2
CrossRef - Hameeda B, Harini G, Rupela OP, Wani SP, Reddy G. Growth promotion of maize by phosphate-solubilizing bacteria isolated from composts and macrofauna. Microbiological research. 2008; 163(2), 234-242. https://doi.org/10.1016/j.micres.2006.05.009
CrossRef - Figueroa-Brambila KM, Escalante-Beltrán A, Montoya-Martínez AC, Díaz-Rodríguez AM, López-Montoya ND, Parra-Cota FI, de Los Santos-Villalobos S. Bacillus cabrialesii: five years of research on a novel species of biological control and plant growth-promoting bacteria. Plants. 2023; 12(13) : 2419. https://doi.org/10.3390/plants12132419
CrossRef - Wang B, Peng H, Wu W, Yang B, Chen Y, Xu F, Peng Y, Qin Y, Fu P, Lu J. Genomic insights into biocontrol potential of Bacillus stercoris 3 Biotech. 2021 Nov;11(11):458. doi: 10.1007/s13205-021-03000-6.
CrossRef - Gatson JW, Benz BF, Chandrasekaran C, Satomi M, Venkateswaran K, Hart ME. Bacillus tequilensis nov., isolated from a 2000-year-old Mexican shaft-tomb, is closely related to Bacillus subtilis. J. Syst. Evol. Microbiol. 2006; 56(7), 1475-1484.
CrossRef - Singh RP, Jha PN. The PGPR Stenotrophomonas maltophilia SBP-9 augments resistance against biotic and abiotic stress in wheat plants. Frontiers in microbiology. 2017.8: p.1945. https://doi.org/10.3389/fmicb.2017.01945
CrossRef - Gao J, Li BY, Wang HH, Liu ZQ.Pseudomonas hunanensis nov., Isolated from Soil Subjected to Long-Term Manganese Pollution. Curr Microbiol 69, 19–24 (2014). https://doi.org/10.1007/s00284-014-0545-4
CrossRef


0 Replies to “Unravelling Promising Diazotrophic Microbiota of Rauvolfia tetraphylla L. with Existing Conventional Nitrogen-Fixing Microbial Partners”