Introduction
Cauliflower (Brassica oleracea var. botrytis L.) is a major crop of the Brassicaceae family, valued for its edible white curd.¹ Native to the Mediterranean region, it has gained global popularity due to its rich nutritional profile, comprising essential vitamins, minerals, and antioxidants. Its ability to thrive under diverse climatic conditions has facilitated widespread cultivation across temperate and tropical regions.² In India, cauliflower plays a significant economic role, being one of the most extensively cultivated vegetables. The country’s varied agro-climatic zones enable year-round production, fulfilling both domestic consumption and export demands.³ However, to meet the increasing demand and overcome various biotic and abiotic stresses, it is imperative to improve both the yield and quality of cauliflower. This requires an in-depth understanding of the genetic variability present in the crop.⁴
Genetic variability refers to the variation in genetic composition among individuals within a species and serves as a cornerstone of plant breeding. Assessing this variability is crucial as it determines the scope for selecting and improving elite genotypes.⁵ A broader genetic base enhances the potential to develop varieties that are high-yielding, nutritionally superior, and resilient to stresses.⁶ To effectively utilize this variability, it is essential to understand the interrelationships among various agronomic traits. Correlation analysis facilitates the identification of the strength and direction of associations between traits.⁷ However, correlation alone does not imply causation.⁸ To overcome this limitation, path coefficient analysis is employed, which partitions correlation coefficients into direct and indirect effects, thereby clarifying the actual impact of individual traits on yield.⁹ This approach helps differentiate traits that influence yield directly from those that act indirectly through other characters.¹⁰ Such analytical tools are invaluable to plant breeders, offering a targeted framework for selecting traits that significantly enhance yield and quality, thus streamlining cultivar development.
In this context, the present study was designed to evaluate 12 cauliflower genotypes obtained from the All India Coordinated Research Project (AICRP) on Vegetables. These genotypes represent a wide genetic spectrum, providing an ideal resource to assess genetic variability and analyze inter-trait associations. By employing correlation and path coefficient analyses, the study seeks to identify key traits contributing to yield and quality, offering insights that can guide the development of improved cauliflower varieties tailored to agronomic requirements and market expectations.
Materials and Methods
During the 2022–2023 growing season, a field experiment was conducted at the Experimental Research Farm of Horticulture, School of Agricultural Sciences, Nagaland University, Medziphema Campus, Nagaland. The primary objective was to evaluate 12 cauliflower (Brassica oleracea var. botrytis L.) genotypes obtained from the All India Coordinated Research Project (AICRP) on Vegetables. The experiment was arranged in a Randomized Block Design (RBD) with three replications to ensure statistical validity and reduce environmental variability.¹¹ Recommended agronomic practices were meticulously followed throughout the crop cycle to maintain ideal conditions for plant growth and yield performance.
A total of eleven quantitative traits were recorded for phenotypic evaluation: plant height (cm), stalk length (cm), number of leaves per plant, plant spread (cm), curd diameter (cm), curd size (cm²), gross curd weight (g), net curd weight (g), yield per plot (kg), curd compactness, and ascorbic acid content (mg/100g). These measurements were taken in accordance with the standard guidelines set by the Protection of Plant Varieties and Farmers’ Rights Authority (PPVFRA).
The collected data were analyzed using OPSTAT, an open-source statistical software, to ensure precise and dependable results.¹² Analysis of variance (ANOVA) and genetic variability parameters were computed using the formulae proposed earlier.¹³ Genotypic, phenotypic, and environmental coefficients of variation were estimated following the methods outlined by Lnu et.al.¹⁴ Heritability and genetic advance were determined based on the approach described by Sarker et al.¹⁵ Correlation coefficients were calculated using standard procedures,¹⁶ while path coefficient analysis was conducted as per the methodology suggested by Chatterjee et al.¹⁷
Results
Analysis of variance
The analysis of variance (ANOVA) revealed highly significant differences (P < 0.01) among the 12 cauliflower genotypes for the agronomic traits evaluated (Table 1). This indicates the presence of substantial genetic variation within the population, highlighting the potential for identifying and selecting superior genotypes for use in breeding programs.
Table 1: Analysis of variance for eleven agronomic traits among twelve cauliflower genotypes
| Character | Mean Squares | CD at 5 % | CV (%) | ||
| Replication df (2) | Treatments df (11) | Error df (22) | |||
| Yield per plot (kg) | 1.262 | 10.204** | 0.401 | 1.073 | 12.666 |
| Plant height (cm) | 4.044 | 59.564** | 1.605 | 2.145 | 3.466 |
| Plant spread (cm) | 1.909 | 166.862** | 1.417 | 2.016 | 2.757 |
| Stalk length (cm) | 0.841 | 8.271** | 0.335 | 0.98 | 6.266 |
| Number of leaves per plant | 0.249 | 16.073** | 1.431 | 2.026 | 7.936 |
| Curd diameter (cm) | 0.409 | 3.828** | 0.655 | 1.37 | 7.216 |
| Curd size (cm2) | 502.59 | 755.164** | 66.039 | 13.761 | 11.174 |
| Gross curd weight (g) | 238.256 | 16062.433** | 802.095 | 47.957 | 4.934 |
| Net curd weight (g) | 5,644.11 | 40924.093** | 1,776.12 | 71.363 | 13.443 |
| Curd compactness | 21.56 | 315.039** | 31.811 | 9.55 | 12.965 |
| Ascorbic acid (mg/100g) | 0.104 | 9.304** | 0.392 | 1.061 | 3.724 |
| ** = Significant at 1 % and * = Significant at 5 % level of significance | |||||
Estimates of genetic variability among twelve cauliflower genotypes
Assessing genetic variability is crucial for determining the extent of differences among genotypes, which facilitates effective selection and crop improvement. The genetic variability parameters for twelve agronomic traits in cauliflower genotypes are summarized in Table 2.
The results revealed that the phenotypic coefficient of variation (PCV) was consistently higher than the genotypic coefficient of variation (GCV) for all traits, indicating the influence of environmental factors on the expression of these traits. Among the traits studied, PCV ranged from 10.90% for ascorbic acid content to 38.84% for net curd weight, while GCV ranged from 9.17% for curd diameter to 36.44% for net curd weight. Traits such as yield per plot, net curd weight, curd compactness, and curd size exhibited high genetic variability, suggesting they hold strong potential for improvement through selection. Traits like plant height, plant spread, stalk length, number of leaves per plant, curd diameter, and gross curd weight showed moderate variability, indicating that selection may still be effective, though environmental influences may play a role. Ascorbic acid content was the only trait that exhibited low genetic variability. All the evaluated traits recorded high heritability estimates (>60%), suggesting strong genetic control and a favorable response to selection. Furthermore, all traits except curd diameter displayed high genetic advance as a percentage of the mean, indicating the predominance of additive gene action. This reinforces their suitability for genetic improvement through direct selection. All investigated traits exhibited both high heritability and high genetic advance as a percentage of the mean except crude diameter. Which intricate these traits controlled by additive gene action and selection among genotypes might be reward.
Table 2: Estimation of genetic parameters for eleven agronomic traits among twelve cauliflower genotypes
| Response Variable | SED | Heritability | GCV | PCV | Gen-Advance | Gen-Adv % Means |
| Yield per plot (kg) | 1.073 | 89.06 | 36.139 | 38.294 | 3.514 | 70.255 |
| Plant height (cm) | 2.145 | 92.331 | 12.026 | 12.516 | 8.7 | 23.805 |
| Plant spread (cm) | 2.016 | 97.495 | 17.198 | 17.417 | 15.105 | 34.981 |
| Stalk length (cm) | 0.98 | 88.77 | 17.616 | 18.698 | 3.157 | 34.192 |
| Number of leaves per plant | 2.026 | 77.327 | 14.656 | 16.667 | 4.002 | 26.55 |
| Curd diameter (cm) | 1.37 | 61.76 | 9.171 | 11.669 | 1.665 | 14.846 |
| Curd size (cm2) | 13.761 | 77.671 | 20.841 | 23.647 | 27.516 | 37.836 |
| Gross curd weight (g) | 47.957 | 86.379 | 12.425 | 13.369 | 136.551 | 23.789 |
| Net curd weight (g) | 71.363 | 88.02 | 36.439 | 38.84 | 220.776 | 70.425 |
| Curd compactness | 9.55 | 74.797 | 22.336 | 25.826 | 17.311 | 39.793 |
| Ascorbic acid (mg/100g) | 1.061 | 88.33 | 10.245 | 10.9 | 3.337 | 19.835 |
Correlation
The genotypic and phenotypic correlation coefficients of twelve agronomic traits with yield per plot are presented in Table 3. The results indicate that most traits show a positive and significant correlation with yield at both the genotypic and phenotypic levels, highlighting their importance in improving productivity. Traits such as plant height, plant spread, stalk length, curd diameter, gross curd weight, net curd weight, curd compactness, and ascorbic acid content demonstrated strong and significant associations with yield per plot. These findings suggest that selection based on these traits could effectively enhance cauliflower yield. Furthermore, the genotypic correlation coefficients were generally higher than the phenotypic ones, indicating a strong underlying genetic association among these traits with relatively lower environmental influence. This emphasizes the potential for successful selection and genetic improvement based on these key agronomic characteristics.
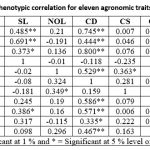 |
Table 3: Estimation of genotypic and phenotypic correlation for eleven agronomic traits among twelve cauliflower genotypes |
Path analysis
Path coefficient analysis was conducted at the genotypic level, and the results are summarized in Table 4. This analysis helps to dissect the direct and indirect effects of various agronomic traits on yield per plot, offering valuable insights into the most influential traits for selection. The findings revealed that traits such as stalk length, number of leaves per plant, curd diameter, net curd weight, and curd compactness had a direct positive effect on yield. Among these, net curd weight emerged as the most influential trait, indicating its strong contribution to yield enhancement. The low residual effect observed suggests that the traits included in the analysis account for nearly all the variation in yield per plot, leaving very little unexplained. This emphasizes the effectiveness of these traits in selection strategies and their potential to significantly improve cauliflower productivity.
Table 4: Estimation of genotypic path coefficient analysis for eleven agronomic traits among twelve cauliflower genotypes
| PH | PS | SL | NOL | CD | CS | GCW | NCW | CC | ACC | YPP (rg) | |
| PH | -0.009 | -0.021 | 0.017 | -0.002 | 0.028 | -0.002 | -0.03 | 0.756 | 0.05 | -0.003 | 0.786** |
| PS | -0.007 | -0.026 | 0.009 | 0.001 | 0.05 | -0.004 | -0.026 | 0.952 | 0.043 | -0.005 | 0.988** |
| SL | -0.006 | -0.01 | 0.024 | 0.000 | -0.007 | 0.011 | -0.011 | 0.462 | 0.022 | -0.001 | 0.485** |
| NOL | 0.002 | -0.004 | 0.000 | 0.010 | 0.033 | -0.017 | -0.009 | 0.205 | -0.008 | -0.002 | 0.210 |
| CD | -0.004 | -0.021 | -0.003 | 0.005 | 0.063 | -0.013 | -0.033 | 0.72 | 0.035 | -0.004 | 0.745** |
| CS | 0.000 | -0.002 | -0.006 | 0.004 | 0.018 | -0.047 | -0.001 | 0.022 | 0.022 | -0.001 | 0.007 |
| GCW | -0.006 | -0.015 | 0.006 | 0.002 | 0.048 | -0.002 | -0.044 | 0.544 | 0.038 | -0.002 | 0.569** |
| NCW | -0.007 | -0.026 | 0.012 | 0.002 | 0.047 | -0.001 | -0.025 | 0.96 | 0.043 | -0.005 | 0.999 |
| CC | -0.008 | -0.019 | 0.01 | -0.001 | 0.038 | -0.018 | -0.029 | 0.725 | 0.057 | -0.005 | 0.750** |
| ACC | -0.003 | -0.018 | 0.003 | 0.003 | 0.036 | -0.01 | -0.016 | 0.764 | 0.038 | -0.007 | 0.790** |
Discussion
This study offers a comprehensive evaluation of genetic variability, heritability, correlation, and path coefficient analysis for key agronomic traits in cauliflower, providing valuable insights for crop improvement programs.¹⁶ Understanding the magnitude of genetic variation and the interrelationships among traits is crucial for identifying superior genotypes with desirable characteristics. The findings lay a solid foundation for future cauliflower breeding efforts aimed at enhancing yield potential and stability.¹⁷ Additionally, this research contributes to a broader understanding of the genetic parameters that influence trait expression, supporting applications in marker-assisted selection and advanced breeding strategies.¹⁸
The results are consistent with earlier studies that reported significant genetic variation among Brassica genotypes. The highly significant differences (P < 0.01) observed in ANOVA among the 12 cauliflower genotypes confirm substantial genetic variability, which is critical for effective selection.¹² Similar trends have been documented in cabbage, reinforcing the potential for genetic improvement.¹²
In this study, the phenotypic coefficient of variation (PCV) consistently exceeded the genotypic coefficient of variation (GCV) across all traits, indicating a notable influence of environmental factors on trait expression.³ This pattern is consistent with findings in cabbage and other Brassica crops, where environmental variability impacts quantitative traits such as yield and nutritional quality.¹⁹ High genetic variability was recorded for traits like yield per plot, net curd weight, curd compactness, and curd size, while moderate variability was observed in plant height, plant spread, stalk length, curd diameter, and gross curd weight. In contrast, ascorbic acid content exhibited low genetic variability.²⁰˒²¹
All evaluated traits showed high heritability estimates (>60%), suggesting strong genetic control.⁴ Traits with both high heritability and high genetic advance as a percentage of the mean indicate additive gene action.⁵ However, curd diameter, despite its high heritability, showed lower genetic advance, implying the involvement of dominance or epistatic effects, which favors heterosis breeding.¹³ Similar observations were reported in cabbage for traits such as head weight and compactness.⁵˒⁶
Positive and significant genotypic correlations with yield were found for traits including plant height,⁷ plant spread,²¹ curd diameter,⁸ curd compactness,⁷ and ascorbic acid content.²³ Genotypic correlations were generally stronger than phenotypic ones, indicating robust genetic associations.⁴˒⁹˒²¹ Path coefficient analysis revealed that net curd weight had the most substantial direct effect on yield,²¹ followed by curd diameter and curd compactness, with plant spread and stalk length also contributing positively.¹⁸ The minimal residual effect observed confirmed the effectiveness of these traits in improving yield.³
Conclusion
The present study revealed significant genetic variation among 12 cauliflower genotypes, highlighting their potential for yield improvement. Characters such as net curd weight, curd compactness, and curd size exhibited high genetic variability, heritability, and genetic advance, indicating their suitability for selection-based breeding programs. The consistently higher PCV than GCV across all traits suggests environmental influence, but the strong genotypic correlations and direct effects on yield confirm the predominance of genetic control. Net curd weight showed the highest direct positive effect on yield, making it the most critical trait for selection. The low residual effect in path analysis indicates that the studied traits explain nearly all the yield variation, ensuring selection efficiency. These findings underscore the importance of net curd weight, curd compactness, and plant spread as primary selection criteria for genetic enhancement in cauliflower breeding programs. Future research should integrate molecular breeding and genotype-by-environment studies to further refine selection strategies and enhance cauliflower productivity.
Acknowledgement
I sincerely thank S. P. Kanaujia for providing valuable guidance and research materials, which greatly contributed to the successful completion of this study.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
All authors declared there is no conflict of interest on this study.
Data Availability Statement
This statement does not apply to this article
Ethics Statement
This research did not involve human participants, animal subjects, or any
material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Permission to reproduce material from other sources
Not Applicable
Author Contributions
All authors contributed significantly in the preparation of the manuscript. All authors have read and approved the final version of the manuscript.
Satya Prakash Kanaujia: Conceptualization, experimental design, and data interpretation.
Rishabh Kumar Singh: Conducted the field experiments and collected the necessary data.
Ashwini Ananda: Statistical analysis and data visualization.
Moakala Changkiri: Literature review and contributed to manuscript writing.
Ravi Shankar: Laboratory analysis and validation of results.
Hanuman Prasad Chaturvedi: Reviewed and edited the manuscript.
Rajat Rajput: Formatting and references, ensuring adherence to journal guidelines.
Ajeet Kumar: Data collection and analysis, providing valuable inputs to the study.
References
- Sharma P., Singh S. P., Dogra B. S., Rani R., Verma A. Assessment of genetic variability, correlation and path analysis studies in cabbage (Brassica oleracea) under low hill conditions of Himachal Pradesh. Indian J Agric Sci, 2023;93(12):1375-1379.
CrossRef - Özer M. Ö., Kar H., Bekar N. K., Doğru S. M., Beşirli G., Sönmez İ. Correlation, genetic variability, heritability and genetic advance for some morphological traits in red cabbage lines (Brassica oleracea var. capitata subvar. rubra). J Agric Fac Gaziosmanpaşa Univ (JAFAG), 2023;40(2):58-65.
- Ghiffar H. M., Rostaman R., Nurchasanah S., Syaukany R. F., Manoworn C. Performance of five introduced cabbage varieties (Brassica oleracea var. capitata L.) planted at the highland of Pemalang Regency. E3S Web Conf, 2025;609:06003.
CrossRef - Tanni S. T., Islam M. N., Sultana S., Harun-Ur-Rashid M., Rahim M. A. Genetic variability studies of “flowering Chinese cabbage (Brassica rapa subsp. Chinensis” in Bangladesh. Adv Zool Bot; 2023,11(5):384-391.
CrossRef - Rana N., Sharma A., Kumari V., Lata H., Kaur M., Thakur A. Assessment of genetic variability and character association in mid-late/late cauliflower genotypes. Electron J Plant Breed; 2024,15(1):70-79.
CrossRef - Meena M. L., Ram R. B., Lata R., Sharma S. R. Genetic variability for morphological and qualitative traits in cabbage (Brassica oleracea var. capitata L.). Indian J Plant Genet Resour, 2012;25(3):270-273.
- Priyanka, Singh A., Raghav M., Kumar V. Correlation analysis of leafy mustard (Brassica juncea var. rugosa) genotypes for morphological and nutraceutical attributes under Tarai condition of Uttarakhand, India.
- Verma A., Singh Y. Correlation between downy mildew resistance and yield-related traits in cauliflower (Brassica oleracea L. var. botrytis) under sub-temperate conditions of North Western Himalayas. Vegetos, 2024;1-8.
CrossRef - Panse V. G., Sukhatme P. V. Statistical method for agricultural workers. ICAR, New Delhi, 1978.
- Burton G. W., DeVane E. H. Estimating heritability in tall fescue (Festuca arundinacea) from replicated clonal material. Agron J, 1953;45(10):478-481.
CrossRef - Allard R. W. Principles of plant breeding. John Wiley and Sons, New York, 1960;485.
- Al-Jibouri H. W., Miller P. A., Robinson H. F. Genotypic and environmental variances and co-variances in an upland cotton cross of inter-specific origin. Agron J, 1958;50:633-636.
CrossRef - Dewey D. R., Lu K. H. A correlation and path coefficient analysis components of crest wheatgrass seed production. Agron J, 1959;51:515-518.
CrossRef - Lnu R., Dhall R. K., Manchanda P., Kumari P., Singathiya P. Study on characters associations and path coefficient analysis for yield and quality traits of parthenocarpic cucumber genotypes in poly-net house conditions. Discover Plants, 2025;2(1):1-15.
CrossRef - Sarker U., Azam M. G., Talukder M. Z. A. Genetic variation in mineral profiles, yield-contributing agronomic traits, and foliage yield of stem amaranth. Genetika, 2022;54(1):91-108.
CrossRef - Singh S. R., Rajan S., Veena G. L., Kumar D., Muralidhara B. M., Srivastava K. K. Genetic variability, character association and path analysis for yield and yield attributing traits in lettuce (Lactuca sativa). Indian J Agric Sci, 2020;90(11):2071-2075.
CrossRef - Chatterjee S., Aralikatti O., Sharma S., Mukherjee D., Patil S., Kanwar H. S., Choudhuri P. Studies of genetic variability, heritability and genetic gain for some important horticultural traits in cauliflower (Brassica oleracea var. botrytis L.). Int J Curr Microbiol Appl Sci, 2018;7(4):82-92.
CrossRef - Kindo S. S., Singh D. Varietal evaluation of cauliflower (Brassica oleracea L. var. botrytis) under agro-climatic condition of Allahabad. Int J Pure Appl Biosci, 2018;6(1):672-677.
CrossRef - Kumar V., Singh D. K., Panchbhaiya A. Genetic variability studies in mid-season cauliflower (Brassica oleracea var. botrytis L.). Bull Environ Pharmacol Life Sci, 2017;6(9):99-104.
- Kumar L., Gayen R., Singh J., Mehat N. Estimation of genetic variability, heritability and genetic advance in Indian cauliflower (Brassica oleracea var. botrytis L.). J Pharmacogn Phytochem, 2019;8(6):233-235.
- Lakshmi T. S., Kanaujia S. P., Jha A. Evaluation of cauliflower genotypes for growth, yield and quality traits under foothill conditions of Nagaland. Prog Hortic, 2022;54(1):112-115.
CrossRef - Verma A., Singh Y. Inheritance of downy mildew resistance and its relationship with biochemical traits in cauliflower (Brassica oleracea L. var. botrytis). Crop Prot, 2018;106:132-138.
CrossRef - Vanlalneihi B., Saha P., Srivastava M. Assessment of genetic variability and character association for yield and its contributing components in mid-maturing Indian cauliflower. Int J Curr Microbiol Appl Sci, 2017;6(11):2907-2913.
CrossRef

