Introduction
Tree tomato (Solanum betaceum Cav.) belongs to the Solanaceae family.1 It is considered a sub-tropical tree and is generally grown in well drained soils between 1000 m to 3,000 m altitude.2 It is native to the Andes of Ecuador, Chile, Peru and Bolivia. In Kenya, it is mainly cultivated in Makueni, Nandi, Baringo, Laikipia, Nyeri, Tharaka Nithi, Murang’a, Nakuru, Narok, Kiambu, Kirinyaga, Meru, Elgeyo Marakwet, Uasin Gishu, and Nyandarua counties.3 It is highly sought by dieticians because of the high amounts of vitamins A and C, and minerals it contains, and due to it being low in calories.4 Tree tomato fruit extracts have also been observed to confer therapeutic health benefits such as helping manage cholesterol levels in obese individuals, and exhibiting cancer prevention and treatment in liver and breast cancer cells lines.5,6
The main constraints facing tree tomato cultivation in Kenya are diseases.7 Studies have observed viruses to be the most significant phytosanitary constraint to tree tomato production.8,9 Prior to this study, only potato virus Y (PVY) and tomato mild mottle virus (TMMoV) have been found to infect tree tomato in Kenya.10–12
Potato virus Y is a member of the Potyvirus genus, from the Potyviridae family. It is spread by aphids in a non-persistent fashion, and has a large host range, about 405 species from 31 families, with most of these, 287 species, coming from the Solanaceae family.13,14 Because of its worldwide distribution and wide host range, it is considered to be among the top ten most important plant viruses affecting crops, mainly potato, tomato, tobacco, pepper, and eggplant.13,14 Potato virus Y has previously been found to cause disease in tree tomato in other countries like Colombia and New Zealand.9,15 Solanaceous perennials like tree tomato are considered to be natural PVY reservoirs and ideal models that foster virus evolutions.16 Recently, three novel PVY recombinant strains reported in Ecuador are thought have co-evolved in tree tomato. In Kenya, PVY was first reported to infect tree tomato using traditional diagnostic techniques―dilution end point, longevity in vitro, electron microscopy, thermal inactivation point and symptomatology.10 To the best of our knowledge, there is no record of any study in Kenya that has established the presence of PVY in tree tomato using modern molecular techniques.
This survey was conceptualised to determine the presence, and distribution of PVY in tree tomato growing counties in Kenya, across different agro-ecological zones (AEZ), using modern molecular techniques, including RT-PCR, and Sanger sequencing. This study intended to provide important baseline data for the development of effective virus management strategies in tree tomato production.
Materials and methods
Survey and sample collection
Between December 2018 and August 2019, two surveys were conducted in nine tree tomato growing counties within the three AEZ, i.e., upper highland (UH) – 2450m – 3050m, lower highland (LH) – 1850m – 2450m and midland (ML) – 1500m to 1850m.17 The counties included Baringo, Nakuru, Nandi, Elgeyo Marakwet, Meru, Tharaka Nithi, Embu, Machakos, and Nairobi. Disease severity was determined visually and using a 0 to 5 ordinal rating scale adapted from Murphy18 as follows: 0=Symptomless; 1=mottling, yellowing on young leaves, mild mosaic; 2=obvious mottling, mosaic, yellowing on leaves from at least one of the main stems, vein clearing and banding; 3=obvious vein banding and clearing on leaves over of the entire plant, mottling, mosaic; 4=obvious vein banding and clearing, mosaic, mottling, severe stunting, and leaf malformation; and 5=obvious vein banding and clearing, mosaic, mottling, severe stunting, leaf malformation and necrosis. The mean severity per county AEZ were used for comparison. Disease incidence was expressed as a percentage of all plants examined per AEZ in a county. Disease prevalence was determined by the percentage number of fields with virus associated symptoms per county AEZ. Leaf samples from each farm were collected and preserved by pressing in paper towels followed by desiccation in zip-lock bags using silica gel. The samples were taken to the molecular laboratory at KALRO Biotechnology Centre for virus detection.
Sample preparation and RT-PCR analysis
Total RNA was extracted from a 1 cm2 portion of each leaf sample using the CTAB protocol detailed by Gambino19 with slight modifications. A Nanodrop spectrophotometer was used to determine the concentration of RNA, followed by dilution to 0.5 µg/µl. A 10 µl RT-PCR reaction mix was prepared following the manufacturer’s protocol: 5 µl of the reaction mix, 2.2 µl of sterile water, 0.4 µl of the SuperScript™ III RT and Platinum™ Taq mix, 0.2 µl of each forward and reverse primer at 10 µM concentration, and lastly 2 µl of the RNA template. A pair of universal primers for PVY, F3 – CGTTGAAACCAATCGTTGAGAA and B3 – GACATCCTCGGTGGTGTG were used.20 The reaction was done as follows: reverse transcription was done for 30 min at 50°C; initial denaturation for 2 min at 94°C; cyclic steps included denaturation for 20 s at 94°C, annealing for 20 s at 58°C, and extension for 30 s at 68°C. A total of 35 cycles were conducted followed by a final extension for 5 min at 68°C. A 1% agarose gel was prepared to view the results. One confirmed positive sample per farm was subjected to Sanger sequencing.
Data analysis
One-way analysis of variance was used to assess disease severity and incidence across counties in the three different AEZ with significance determined at P < 0.01. Where significant differences were detected, the HSD (Honestly Significant Difference) Tukey-Kramer post hoc test was applied for comparing significant differences in the means of prevalence, severity, and incidence scores.
Virus incidence was calculated using percentage number of samples in a farm which tested positive for PVY. Due to the absence of PVY in one of the AEZ, UH, a student t-test at 95% confidence level was used to compare the mean virus incidence. The distribution of PVY was mapped based on the counties from which samples were collected. Multiple sequence alignment was done using Clustal Omega, and a maximum likelihood tree was inferred using MEGA 11. Recombination on the fourth recombination junction, on which the target region lies, was tested using RDP4.
Results
Survey data
A total of 358 leaf samples were obtained from farms located across the three AEZ: ML, LH and UP. The samples were collected from 26 farms in different counties as follows: two from Meru, three from Machakos, four from Tharaka Nithi, five from Embu, three from Elgeyo Marakwet, one from Nairobi, one from Naivasha and four from Nandi. Of the 26 farms, 10 were from ML, 11 from LH and five from UH. All orchards intercropped tree tomatoes with either tomatoes, tobacco, potatoes, peppers, capsicum, Tea, oranges, beans, banana, sweet potato, macadamia, maize, coffee, apples, cassava, arrow roots, avocado, or kale. The famers identified three constraints to tree tomato production, pests and diseases bad weather and weeds. At the time of the study the farmers claimed to have no access to certified virus-free planting material. All the surveyed crops were in the fruiting stage and nearing harvest. Few farmers practiced pest or disease control, and even among them, disease symptoms persisted.
A range of virus-associated symptoms were observed across the surveyed farms, including vein clearing, crinkling, leaf malformation, necrotic spots, and vein banding. Disease prevalence was 100% across all regions. Disease incidence per county AEZ ranged between 60 to 80%, whereas severity ranged between 2.1 and 3.5. The highest disease severity was recorded in Meru County, whereas the highest disease incidence was recorded in Baringo County (Table 1). A one-way ANOVA revealed a statistically significant difference in the disease severity (p<0.01) among UH, ML and LH. However, the incidence was not statistically significant (Table 1). Subsequent analysis using the Tukey-Kramer test for post hoc analysis at α<0.01 identified a significant difference in severity scores between UH, and LH and ML. No statistically significant difference was detected in severity scores between the LH and the ML.
Table 1: Incidence and Severity scores of virus-associated symptoms in the surveyed counties across different AEZ during 2018-2019 growing season, with ANOVA results, including post hoc comparisons using the HSD Tukey-Kramer test.
| County (AEZ) | Severity | Incidence | |
| Embu (ML) | 2.6 | 70% | |
| Elgeyo Marakwet (UH) | 2.2 | 70% | |
| Tharaka-Nithi (ML) | 3 | 65% | |
| Machakos (LH) | 2.3 | 70% | |
| Nandi (LH) | 2.4 | 75% | |
| Nairobi (ML) | 3.2 | 60% | |
| Nakuru (LH) | 2.1 | 80% | |
| Meru (LH) | 3.5 | 70% | |
| Baringo Farm 23 (LH) | 3.15 | 80% | |
| Baringo Farms 24 and 25 (UH) | 2.4 | 80% | |
| Significance at P<0.01 | 1.406e-32 | 0.35 | |
| Significance | sd | nsd | |
| Severity post-hoc analysis | ML/LH LH/UH UP/ML | ||
| Studentized Range (Q) | 3.46 14.61 16.43 | ||
| HSD (α=0.01) | 4.13 | ||
sd – significant difference at p < 0.01; nsd – no significant difference; ML – midlands; LH – lower highlands; UH – upper highlands.
RT-PCR results
The RT-PCR reactions revealed that PVY was prevalent in all the counties surveyed except Elgeyo Marakwet. Virus incidence per county ranged between 25% and 100%. A total of 12 farms out of 26 farms had samples which tested positive, representing 46% of all farms surveyed (Figure 1).
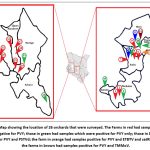 |
Figure 1: Map showing the location of 26 orchards that were surveyed. The Farms in red had samples which tested negative for PVY; those in green had samples which were positive for PVY only; those in blue were positive for PVY and PSTVd; the farm in orange had samples positive for PVY and ETBTV and satRNA-E; and the farms in brown had samples positive for PVY and TMMoV. |
RT-PCR detected PVY in 72 individual samples (Figure 2). Samples from these farms showed various symptoms including leaf mottling, chlorosis, mosaic patterns and leaf malformation. Of these, 36 samples from seven farms were infected by PVY only (gels A-E). The other 36 samples positive for PVY shown in gels F – J were also positive for a second virus, including: TMMoV, PSTVd, and ETBTV and satRNA-E.21–23
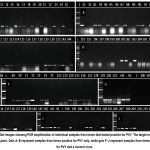 |
Figure 2: Gel images showing PCR amplification of individual samples from farms that tested positive for PVY. The target region was 331 base pairs. Gels A–E represent samples from farms positive for PVY only, while gels F–J represent samples from farms positive for PVY and a second virus. |
Plants in the farms which had a co-infection with PSTVd exhibited similar symptoms to those which were positive for PVY only, but with increased severity.21 Mottling, mild necrosis on the veins, and leaf curling were observed in farms 14 and 19 where all samples had a mixed infection of PVY and TMMoV.23 Farm 11 which had plants with a mixed infection of PVY, and ETBTV and satRNA-E exhibited a more severe combination of symptoms, including leaf curling, malformation and vein clearing.22 There was no farm from the UH which was positive for PVY. An independent two-sample t-test showed no statistically significant difference in mean virus incidence between the ML (mean = 54.22) and LH (mean = 29.05) zones (t = 1.26, p = 0.27, two-tailed).
Nucleotide sequences from representative PVY-positive samples from the 12 farms, ON159312 – ON159323, showed a high level of nucleotide sequence similarity to each other, ranging between 98% to 100, and had a negative Tajima D value of –0.1283. Recombination Detection Program 4 (RDP4) showed no evidence of recombination on the fourth recombinant junction normally present in the CP region. The sequences showed a high percentage identity, 97 to 98% to strain N:O. Likewise, a maximum likelihood phylogenetic tree comparing Kenyan PVY isolates from tree tomato with 24 known PVY strains revealed a close relationship with the recombinant strain N:O (Figure 3).24
 |
Figure 3: A maximum likelihood phylogenetic tree was constructed using MEGA 11. The analysis included 12 nucleotide sequences of PVY isolates from this study (accession numbers ON159312 to ON159323) alongside 24 reference sequences representing known parent and recombinant PVY strains. Carrot virus Y (NC004013) was used as the outgroup. |
Discussion
Detection of PVY in tree tomato using RT-PCR revealed that the virus is prevalent in all counties surveyed except Elgeyo Marakwet, and in all AEZ except the UH zones. The absence of PVY in the UH zones could be as a result of low temperatures experienced in these zones, 16°C to 20°C, which may restrict aphid population which in turn limit the spread of PVY.25
The cases of mixed infection are thought to occur as PVY facilitates the aphid transmission of other viruses as follows: TMMoV by acting as a helper virus, while with PSTVd, ETBTV and satRNA-E, by encapsidation of their virion and genetic material respectively.26–28
The primers used in this study were designed to target a highly conserved region of the CP which enable the detection of both known and emerging PVY isolates. This is reflected by the negative Tajima’s D value and high percentage identity observed. Consequently, the close phylogenetic relationship observed between the Kenyan isolates and strain N:O may be incidental, reflecting sequence similarity in a conserved region rather than overall relatedness. The recombinant strain N:O derives its genome from two parental strains: Eu-N (from the 5′ end to recombinant junction RJ2 at position 2390 bp) and O (from RJ2 to the 3′ end).24 Since the primers target a region near RJ4—where no recombination was observed in this study’s sequences and which is absent in N:O—the resulting amplicons likely represent the O-strain portion of the genome.24 While the primers can reliably detect PVY infections, they are not ideal for differentiating among its various strains. For accurate definition of PVY strains whole genome sequencing and phylogenetic studies are recommended, alongside biological characterisation and symptom expression on its host.29
Understanding the distribution of PVY in tree tomato, a perennial crop, provides starting point for other studies. Future research should investigate the evolution of PVY in tree tomato and other susceptible perennials across different agro-ecological zones (AEZs) in greater depth. This will help to develop effective mitigation strategies to prevent its spread to other solanaceous hosts, particularly tomato and potato, which are economically more important and widely cultivated.
Conclusion
In conclusion, this study has established the presence of PVY using RT-PCR and Sanger sequencing in eight out of nine counties. In total, 12 out of 26 farms had samples which tested positive for PVY, five of which occurred in mixed infection with other viruses, namely ETBTV and satETBTV-E, PSTVd and TMMoV. PVY was prevalent in all AEZ with the exception of UH. These findings establish a critical baseline for virus monitoring and call for integrated management strategies to mitigate the impact of viral diseases on tree tomato production in Kenya.
Acknowledgement
The authors would also like to thank John Irungu for is assistance during the collection of samples, Mary Lechuta her assistance during RNA extraction, and Dr Esther Kimani, and Dr Bramwel Wanjala for their assistance during RT-PCR.
Funding Sources
This study was supported by the Kenya Agricultural and Livestock Research Organisation (KALRO) through funding from USAID grant # AID-615-F-16-00001
Conflict of Interest
The authors do not have any conflict of interest.
Data Availability Statement
The manuscript incorporates all datasets produced or examined throughout this research study.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Permission to reproduce material from other sources
Not Applicable
Author Contributions
Michael Njoroge Kinoga: Conceptualization, Data Collection, Analysis, Methodology, Writing, Project Administration – Original Draft.
Paul Kimemia Kuria: Conceptualization, Data Collection, Analysis, Writing, Project Administration – Review & Editing, Funding Acquisition.
Douglas Watuku Miano: Visualization, Supervision, Project Administration.
Lusike Wasilwa: Funding Acquisition, Resources, Supervision.
References
- Morton JM. The Tree Tomato, or “Tamarillo”, a Fast-Growing, Early-Fruiting Small Tree for Subtropical Climates. Vol 95. Julia F. Morton; 1982.
- Orwa C, Mutua A, Kindt R, Jamnadass R, Anthony S. Agroforestree Database: A Tree Reference and Selection Guide. 4.0. World Agroforestry Centre; 2009.
- Horticulture Validated Report 2019-2020. The Horticultural Crops Directorate; 2020.
- Prohens J, Nuez F. The Tamarillo (Cyphomandra betacea): A Review of a promising small fruit crop. Small Fruits Review. 2001; 1: 43-68.
CrossRef - Abdul Kadir NAA, Rahmat A, Jaafar HZE. Protective effects of tamarillo (Cyphomandra betacea) extract against high fat diet induced obesity in sprague-dawley rats. Journal of Obesity. 2015; 2015: 1-8.
CrossRef - Abdul Mutalib M, Rahmat A, Ali F, Othman F, Ramasamy R. Nutritional compositions and antiproliferative activities of different solvent fractions from ethanol extract of Cyphomandra betacea (tamarillo) fruit. Malaysian Journal of Medical Sciences. 2017; 24: 19-32.
CrossRef - Muriithi C, Matiri F, Kihanda F, Maina D, Kasungo F. Evaluation of potential of tree tomato production and marketing in eastern Kenya for improved nutrition and food security. In: 27th Soil Science Society of East Africa and the 6th African Soil Science Society; 2013.
- Mejía DM, Rodas EI, Patiño LF, González EP. Effect of acibenzolar-s-methyl on virus infection progress caused by potyvirus in tree tomato. Agronomia Colombiana. 2009; 27:87-93.
- Jaramillo M, Gutiérrez PA, Lagos LE, Cotes JM, Marín M. Detection of a complex of viruses in tamarillo (Solanum betaceum) orchards in the Andean region of Colombia. Zerbini MF, ed. Tropical Plant Pathology. 2011; 36: 150-159.
- Mbula BB. Potato Virus Y (PVY) in Irish Potatoes (Solanum Tuberosum) and Tree Tomato (Cyphomandra betaceae) and the Influence of Potato Susceptibility to the Virus and the Aphid Virus-Vector on the Spread of PVY Mosaic in Kenya (PhD). University of Nairobi; 1992.
- Monger WA, Nixon T. Tomato Mild Mottle Virus in Tamarillo in Kenya. Novel Methods, Food and Environment Research Agency; 2010.
- Gekone JM, Too A. Potato Virus Y in Tree Tomato. In: Pest management decision guide: green and yellow list. CAB International; 2016.
CrossRef - Lacomme C, Jacquot E. General Characteristics of Potato virus Y (PVY) and Its Impact on Potato Production: An Overview. In: Potato Virus Y: Biodiversity, Pathogenicity, Epidemiology, and Management. Annual Review of Phytopathology; 2017:1-19.
CrossRef - Scholthof K, S., Adkins, Czosnek H, et al. Top 10 plant viruses in molecular plant pathology. Molecular Plant Pathology. 2011; 12: 938-954.
CrossRef - Eagles RM, Gardner RC, Richard LS, Forster RLS. Incidence and distribution of six viruses infecting tamarillo (Cyphomandra betacea) in New Zealand. New Zealand Journal of Crop and Horticultural Science. 1994; 22: 453-458.
CrossRef - Jridi C, Martin JF, Marie-Jeanne V, Labonne G, Blanc S. Distinct Viral Populations Differentiate and Evolve Independently in a Single Perennial Host Plant. Journal of Virology. 2006; 80: 2349-2357.
CrossRef - Sombroek WG, Braun HMH, van der Pouw BJA. Exploratory Soil Map and Agro-Climatic Zone Map of Kenya, 1980: Scale 1:1,000,000. Kenya Soil Survey; 1982.
- Murphy JF, Zehnder GW, Schuster DJ, Sikora EJ, Polston JE, Kloepper JW. Plant growth-promoting rhizobacterial mediated protection in tomato against Tomato mottle virus. Plant Disease. 2000; 84: 779-784.
CrossRef - Gambino G, Perrone I, Gribaudo I. A Rapid and effective method for RNA extraction from different tissues of grapevine and other woody plants. Phytochemical Analysis. 2008;19.
CrossRef - Przewodowska A, Zacharzewska B, Chołuj J, Treder K. A one-step, real-time reverse transcription loop-mediated isothermal amplification assay to detect Potato Virus Y. American Journal of Potato Research. 2015; 92: 303-311.
CrossRef - Kinoga MN, Kuria PK, Miano DW, Wasilwa LA. First report of Potato spindle tuber viroid infecting tree tomato in Kenya in mixed infection with Potato virus Y. New Disease Reports. 44.
CrossRef - Kinoga MN, Kuria PK, Miano RD D W, Narla, Wasilwa LA. First report of Ethiopian tobacco bushy top virus and its associated satellite RNA in mixed infection with Potato virus Y infecting Solanum betacea in Kenya. New Disease Reports. 44.
CrossRef - Kinoga MN, Kuria PK, Miano DW, et al. Genome characterisation of two complete coding sequences of tomato mild mottle virus from tree tomato and their distribution in Kenya. Journal of Plant Pathology. Published online 2022:15-19.
CrossRef - Green KJ, Brown CJ, Gray SM, Karasev AV. Phylogenetic study of recombinant strains of Potato virus Y. Virology. 2017; 507:40-52.
CrossRef - Radcliffe EB. Insect pests of potato. In: Annual Review of Entomology. 27th ed.; 1982:173-204.
CrossRef - Romanova SA, Volkov YG, Kakareka NN, Pleshakova TI, Kozlovskaya ZN. Potato disease caused by combined infection with potato spindle tuber viroid and Potato virus Y necrotic strain. Russian Agricultural Sciences. 2007; 33: 162-165.
CrossRef - Abraham A, Menzel W, Vetten HJ, Winter S. Analysis of the tomato mild mottle virus genome indicates that it is the most divergent member of the genus Ipomovirus (family Potyviridae). Archives of Virology. 2012; 157: 353-357.
CrossRef - Dombrovsky A, Reingold V, Antignus Y. Ipomovirus – an atypical genus in the family Potyviridae transmitted by whiteflies. Pest Management Science. 2014; 70:1553-1567.
CrossRef - Glais L, Bellstedt DU, Lacomme C. Chapter 3: Diversity, Characterisation and Classification of PVY. In: Potato Virus Y: Biodiversity, Pathogenicity, Epidemiology and Management. Springer Nature; 2017:43-76.
CrossRef

