Introduction
Research on plant disease detection as a topic has started to gain more attention in precision agriculture in light of the need for food security, sustainable agriculture, economic stability, and the growing global need for precision agriculture. Approximately 20 to 40% of agricultural production is lost each year due to pests and pathogen infected crops, which have considerable economic, social, and nutritional effects.1,2 Historically, disease detection has depended on agricultural specialists performing inspections. This is a slow and, to a degree, inaccurate system due to the subjectivity of the inspector. Having a system like this for large scale operations is impractical given the need for rapid and precise disease detection systems to avoid large scale crop losses.3,4 Arguably the greatest downside to inspection systems is the variability based on person to person which speaks to the system. This demonstrates the need for systems to be designed for full automation.
In recent years, the industry has witnessed the growth of Analytical Intelligence (AI) and Deep Learning (DL). One of the major challenges in the field of plant disease identification has been automation, precision, and data driven diagnostics in the field. Convolutional Neural Networks (CNNs) are one of the many architectures in Deep Learning.5,6 They are described as the deep learning framework that learns, and forms hierarchies of the various layers of the raw leaf images. They autonomously understand visual attributes such as lesions, discolorations, and texture variations that are difficult to capture and quantify in traditional image processing. CNNs are embedded in diagnostic tools and systems and support disease pattern recognition and identification for multiple diseases across various crops and manage to retain precision and robustness in disparate settings. Neural networks and deep learning to support automation and provide diagnostic tools in plant disease identification is a game changer. The results are superior to all other traditional methods that integrate image processing, classical machine learning, and DV in terms of consistency, accuracy, scale, and automation.7 Developing segmentation skills, particularly through semantic and instance segmentation, has pinpointed infected and healthy areas, thereby enhancing the classifiers’ localization and classification abilities. Nevertheless, traditional deep learning techniques are still burdened with huge challenges, such as the lack of generalization during varying lighting, different backgrounds, and surrounding clutter.8
To mitigate these challenges, Quantum Computing (QC) is considered as an innovative approach to accelerate and enhance the computational learning process. By employing the principles of superposition, entanglement, and interference, QC obtains capabilities of information representation and processing those classical computers cannot perform efficiently. Quantum algorithms produce extensive parallelism, which results in faster optimization and improved processing of large, high-dimensional datasets. This is possible through the encoding of classical datasets into quantum states. The use of quantum theory in machine learning, or QML, aims to enhance feature representation, accelerate training, and improve classification efficacy within the realms of classical machine learning.9,10 Quantum-enhanced feature spaces are aimed at transforming classical inputs, stretching the feature spaces of computation, and embedding classical inputs within higher-dimensional Hilbert spaces to unlock intricate patterns and correlations necessary for robust generalization.11,12
Recently, Quantum Neural Networks (QNNs) and hybrid quantum–classical approaches have gained attention in image classification, including agricultural imaging and plant pathology.13,14 Quantum algorithms have shown improvements in training convergence, robustness, and interpretability of models, which indicates potential applications in agriculture. Foundational works on image pattern classification in QML within precision agriculture automated the detection of plant pathogens using hybrid architectures which significantly outperformed traditional methods. Enhanced Controlling Parameter-based Humboldt Squid Optimization Algorithm (ECP-HSOA), quantum-inspired optimization algorithms, have been proposed to tune neural networks to reduce training time, and increase the reliability and efficiency of Deep Learning (DL) models.15
Nevertheless, the current use of quantum-enhanced deep learning for plant disease cognition is very nascent and lacking proper development. Most research is lensing on classical DL architectures, while few studies have hybrid quantum–classical models where QNNs are integrated with contemporary networks like Dense Net, attention-based networks, and capsule networks. This highlights the need for more exploration of quantum-enhanced deep learning to build robust, explainable, and scalable systems for real agricultural use that can efficiently detect plant disease.
Modern agriculture increasingly depends on intelligent and automated solutions to address challenges related to crop health, productivity, and global food security. Among these challenges, plant disease detection remains critical, as undiagnosed or late-detected diseases can lead to substantial yield loss, reduced economic viability, and long-term environmental impact. With the rise of artificial intelligence, deep learning has become a powerful tool for analyzing plant images and identifying diseases with high accuracy.
Recent advancements have introduced a new research direction Quantum-Enhanced Deep Learning (QEDL) which explores the integration of quantum computing principles into traditional deep learning workflows. This emerging paradigm offers the potential for faster learning, improved generalization, enhanced computational efficiency, and support for more reliable and scalable decision-making in agriculture. By bridging classical and quantum intelligence, QEDL aims to make modern agricultural systems more accurate, sustainable, and resilient.
This study contributes to the growing body of knowledge by:
Presenting a comprehensive review of both classical deep learning and quantum-enhanced approaches for plant disease detection and outlining their relevance to precision agriculture.
Examining how quantum neural networks and quantum-inspired optimization strategies may strengthen model performance, particularly in large-scale or real-time scenarios.
Identifying research gaps such as hardware limitations, efficient data encoding constraints, and the limited explainability of quantum-driven models, while highlighting future opportunities including explainable quantum AI, quantum-assisted transfer learning, and edge-ready hybrid frameworks.
Literature Review
Recently, growing interest in precision agriculture, sustainable farming, and global food security has propelled the use of Artificial Intelligence (AI) and Deep Learning (DL) technologies into plant disease detection. Plant disease detection through traditional image analysis methods—focusing on texture, color, and shape—worked well on small datasets. However, traditional methods did not scale well. They struggled generalizing over different crops and environments. There was also a lack of illumination, background clutter, and leaf orientation on various images, which made it even harder to accurately detect a diseased plant. All of this highlighted the pressing need to use more sophisticated computing approaches.
In the last ten years, the automated recognition of plant diseases has relied heavily on Deep Learning techniques, particularly Convolutional Neural Networks. CNNs analyze and learn the intricate features embedded within leaf images, such as lesions, color variations, and other morphology alterations. For instance, Inception-based models have effectively been utilized in extracting relevant spatial and textural features for the detection of diseases in the leaves of tomato plants.16 Likewise, models built on AlexNet achieved impressive performance on multi-class classification by distinguishing infected leaves that were visually similar, utilizing the power of deep hierarchal feature analysis.17 The incorporation of feature selection techniques such as multi-level deep entropy-ELM frameworks have made classification in crops like cucumber more accurate by isolating redundant features and focusing on critical distinguishing features.17
Hybrid CNN-based models that integrate feature limitation and dimensionality optimization for range reduction offer improved computational efficiency while preserving accuracy. Machine learning together with digital color imaging has been used for detection of wheat crown rot with specialized surveillance in controlled environments.18 In CNNs, the implementation of dimensionality reduction has effectively trimmed down training times while still maintaining high predictive accuracy. This has been reported in the case of several crop leaves. The value of data acquisition in delivering strong model performance has been demonstrated in the AI-based high-resolution images used in the detection of diseases in guavas. Predictive deep learning models have achieved scalable applicability in multi-class classification that goes beyond the traditional single crop use case, and thus, offers more agricultural utility.
The positive influence of transfer learning and pre-trained models on developing frameworks for plant disease detection is even more pronounced when labeled datasets are scarce. Real-time deployment is made possible by RT-Droid, which maximizes computing efficiency while detection performance remains solid.24 Knowledge cross over and generalization for various plant species are made possible through transfer learning and feature concatenation. Enhanced hybrid models for the detection of disease on olive leaves use several component CNN frameworks, retaining robust accuracy that meets the threshold of computational efficiency. The streamlined NASNetMobile-based LightLeafNet is an example of lightweight networks intended for sped up inference, and more importantly, downsized networks that are beneficial for edge deployment in the IoT environment. On the other hand, the use of improved variants of U-Net for segmentation of disease areas contributes to the interpretability of segmentation and aids in the classification of downstream tasks.19
In deep learning, advanced techniques still face issues like the high cost of computation, long training time, inconsistencies from changing environments, and reliance on large, labeled datasets. This has led to the development of Quantum-Enhanced Deep Learning (QEDL), which combines quantum computing and classical architecture to try to relieve deep learning issues. Quantum-inspired techniques take advantage of superposition, entanglement, and interference to achieve high-dimensional feature mapping, quicker convergence, and better generalization.
Quantum Neural Networks (QNNs): A New Frontier
Quantum Neural Networks replace classical deep learning methods that utilize classical bits since QNNs operate on qubits. QNNs utilize all four features of quantum systems: superposition, entanglement, and quantum interference to facilitate high-dimensional feature embeddings and parallel processing. In plant disease detection, QNNs improve the pattern learning and feature discrimination of quantum state extraction for classical feature extraction on leaf images. Quantum gates, especially the Hadamard and Pauli-X gates, as well as rotation operations, perform nonlinear transformations of differing magnitudes, which enhance various feature spaces. QNNs that use quantum variational circuits maintain lower classifier size which allows them to achieve high levels of accuracy, rendering them suitable for low power edge computing. Many systems that incorporate QNNs have shown improvements on key CNNs in terms of the convergence speed and predictive accuracy. This is establishing significant potential for agricultural real-time use as demonstrated in Kaur et al.20
Quantum-Inspired Optimization Techniques
Deep learning models take prolonged periods to tune many parameters and optimize a model. Quantum-inspired optimization algorithms accelerate convergence and balance the tradeoff between exploration and exploitation. The Enhanced Controlling Parameter-based Humboldt Squid Optimization Algorithm (ECP-HSOA), a model based on dynamics foraging behavior of Humboldt squids, appropriates this approach. ECP-HSOA flexibly modulates the equilibrium between globally exhaustive searches and localized quits, thus averting convergence to suboptimal solutions and enhancing the resolution of local minima. The integration of ECP-HSOA with QNN-augmented frameworks results in improvement over gradient-based optimizers like Adam and SGD with respect to both efficiency and accuracy of plant disease detection models.21
Hybrid Quantum-Enhanced Deep Learning Models
Current research primarily targets hybrid systems that integrate conventional deep learning approaches with quantum boosted components and methods. The most efficient systems utilize DenseNet’s optimal feature reuse and efficient gradient flow alongside attention mechanisms that concentrate on critical areas of the leaves. Capsule Networks that perceived and sustained spatial hierarchies, QNN layers that mapped high-dimensional features, and training optimization with ECP-HSOA also contributed to the optimization of the system. This DenseNet + Attention + Capsule + QNN + ECP-HSOA system exceeds 95% classification accuracy, while standard CNNs reach 90% and DenseNets 92%, and it also reduces the training time by over 30%. These hybrid frameworks not only executed the highly accurate segmentation of infected leaf region and classification of numerous crop diseases, but also performed consistently, regardless of fluctuating atmospheric conditions, demonstrating their suitability for direct integration into smart farming systems.22
Table 1: Performance of Classical and Quantum-Enhanced DL Models
| Approach | Core Architecture | Key Enhancement | Accuracy (%) | Limitation |
| CNN6 | Convolutional layers | None | 88–90 | Overfitting, slow training |
| DenseNet8 | Dense connectivity | Feature reuse | 91–93 | High memory usage. |
| CapsNet + Attention10 |
Capsule routing + attention | Spatial focus | 92–94 | Complex routing mechanism |
| QNN11 | Quantum variational circuits | Quantum encoding | 93–95 | Hardware constraints |
| Hybrid (DenseNet + Attention + Capsule + QNN + ECP-HSOA)12 | Hybrid fusion | Quantum-inspired optimization | 95+ | Implementation complexity |
Table 2: Key Quantum-Enhanced Optimization Strategies for DL
| Optimization Technique | Core Concept | Application in Plant Disease Detection | Advantage |
| ECP-HSOA | Adaptive global-local search inspired by Humboldt squid behavior | Weight initialization and convergence for QNN layers | Faster training, avoids local minima |
| Quantum Variational Circuits | Parametric quantum gates | High-dimensional embedding of leaf features | Improved accuracy with smaller model size |
| Hybrid Dense + Attention + Capsule + QNN | Integration of classical and quantum layers | Multi-crop disease classification | High accuracy, robust under environmental variations |
In the area of detecting plant diseases, considerable progress has been made with conventional deep learning techniques, yet several important issues remain unresolved, highlighting the potential value of quantum enhanced frameworks. Although traditional CNNs, DenseNets, and hybrid CapsNet-Attention systems yield considerable success with accuracies of 88 to 94%, there are challenges of overfitting, memory consumption, convergence time, and environmental instability. Improvements of high-dimensional feature mapping, convergence speed, and generalization by QNNs and quantum-inspired optimizers such as ECP-HSOA are balanced by the complexity of implementation, provision of supportive hardware, and the challenges of encoding high-dimensional image data.23-29 As shown in Table 1, the comparative evaluation of classical, quantum, and hybrid frameworks reveals that the performance of hybrid DenseNet + Attention + Capsule + QNN + ECP-HSOA systems even surpasses 95% accurate performance, while resolving issues with convergence and feature extraction.
Despite advancements in the field, there are still areas that require further exploration: (i) the scalability of quantum systems is constrained by the limited qubit range as well as the noisy quantum hardware; (ii) the research on the efficient and robust quantum encoding of high-dimensional plant images remains frontier; (iii) the hybrid classical quantum models, and integration of attention and capsule networks are areas that hybrid models are yet to be evaluated on; and (iv) the quantum-enhanced functionalities of features that predict diseases and their explainability is still a growing field of research. In Table 2, which summarizes quantum-enhanced optimization strategies, the ECP-HSOA and quantum variational circuits offer numerous advantages, such as improved weight initialization, high-dimensional feature embedding, and training efficiency, yet the challenge of deploying these systems on edge devices for real-time responsive agriculture remains unmet. Closing these will offer high-accuracy quantum-enhanced deep learning frameworks for multi-crop plant disease detection, which will help transition the technology from cutting-edge laboratory research to real-world flexible agriculture.
Methodology
This survey follows a structured review methodology aimed at examining the progression of plant disease detection techniques from classical deep learning models to quantum-enhanced and hybrid quantum–classical approaches. To capture a comprehensive picture of current advancements, a broad literature search was conducted across major academic databases such as IEEE Xplore, ScienceDirect, SpringerLink, ACM Digital Library, arXiv, and Google Scholar. The search used combinations of keywords related to plant disease detection, deep learning architectures, quantum neural networks, and quantum-inspired optimization methods, focusing on studies published between 2015 and 2025.
Relevant research studies were selected using predefined inclusion and exclusion criteria to ensure quality and alignment with the survey objectives. Papers were included if they presented classical deep learning models, quantum or quantum-inspired approaches, or hybrid architectures for image-based plant disease identification. Publications that lacked methodological clarity, experimental evaluation, or relevance to precision agriculture were excluded. For each selected study, essential methodological information such as model architectures, datasets, preprocessing strategies, optimization techniques, evaluation metrics, and identified challenges was systematically extracted to support comparative analysis.
The collected studies were then analyzed thematically to understand how classical deep learning methods have evolved and how quantum principles are beginning to reshape this research domain. Classical approaches were reviewed in terms of their performance, computational requirements, and limitations, while quantum and hybrid models were examined for their potential to enhance generalization, training efficiency, and real-time applicability. Through this comparative evaluation, the survey synthesizes key trends, identifies persistent gaps such as hardware constraints and data encoding challenges, and highlights promising directions for future research. This structured approach ensures that the findings presented in this work are coherent, comprehensive, and reflective of the current state of the field.
Incorporating capsule networks and attention mechanisms into classical models helps address some of these challenges. While capsule networks help maintain spatial hierarchies and relationships between features, attention mechanisms aid the networks in focusing on the disease-affected regions, thus enhancing the classification accuracy. This is particularly useful and important for leaf morphologies and rotation invariant disease detection. In addition, DenseNet architectures improve gradient flow and feature reuse, which helps to mitigate the risk of deeper networks suffering from vanishing gradients. Advanced pre-processing techniques such as feature reduction and image segmentation aid in isolating infected regions which reduces background noise and enhances interpretability of the model overall.
Advanced methodologies include QNNs, which incorporates the use of qubits instead of classical binary bits in the DL paradigm. QNNs provide enhanced parallel processing by exploiting superposition, entanglement and interference, along with high-dimensional feature mapping.19,20 Quantum feature mapping of classical images allows most image features to be quantum-encoded, providing multiple richer embeddings. Non-linear transformations of features through Hadamard, Pauli-X and rotation gates, along with quantum variation circuits, which are designed to economize models, enrich the feature space without loss of accuracy. Quantum and classical models are designed for greater computational efficiency with classical feature extractors with quantum layers.20,21
The use of quantum-inspired optimization, specifically the Enhanced Controlling Parameter-based Humboldt Squid Optimization Algorithm (ECP-HSOA), incorporates advanced methodologies. Under parameter tuning, dynamic behavior of Humboldt squid provides a balance between global exploration and local exploitation.16,29 Within DL training, ECP-HSOA improves weight initialization and decreases the time to learn relative to gradient-based optimizers. The synergistic use of quantum-based optimal and architecture leads to hybrid frameworks for superior performance in plant disease detection.
The last part of the methodology is the assessment of hybrid frameworks that combine DenseNet for feature reuse, attention mechanisms for region focus, capsule networks for hierarchical spatial structure preservation, quantum layers for high-dimensional embedding, and ECP-HSOA for optimized training.21–23 These hybrid architectures have shown the ability to accurately segment diseased regions, classify diseases with a high degree of precision, and perform reliably under varying conditions of the environment. The survey thus outlines the evolution of models in their entirety, from classical CNNs to attention-augmented networks, to quantum-enhanced architectures, and hybrid pipelines.
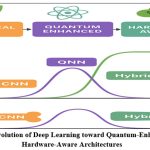 |
Figure 1: Evolution of Deep Learning toward Quantum-Enhanced and Hardware-Aware Architectures |
The figure-1 depicts the various phases in the development of deep learning (DL) paradigms and the transformation from the classical to quantum and hardware-aware paradigms. The upper part shows the evolution path that starts with the Classical Deep Learning (DL) models like CNNs, moves to the Quantum-Enhanced models that utilize the principle of superposition and entanglement through Quantum Neural Networks (QNNs) and finally ends with the Hardware-Aware models that are optimized for hybrid quantum–classical computing deployment. Improvement in model efficiency, convergence, and generalization due to hybrid quantum-classical integration is shown in the CNN, QNN, and Hybrid architectures performance characteristics comparison in the middle and lower plots.
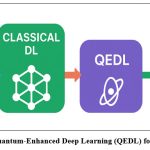 |
Figure 2: Overview of Quantum-Enhanced Deep Learning (QEDL) for Plant Disease Detection |
Figure 2 shows the phased approach of plant disease detection working from the traditional manual inspection, going through the classical Deep Learning (DL) methods, and finally reaching Quantum-Enhanced Deep Learning (QEDL). The stages in the workflows are color coded where the workflows in green are the conventional and DL based methods, purple illustrates the quantum integration, and the steps in teal are the outcomes of the performance. The flow illustrates the integration of quantum principles, with advanced DL frameworks, increases accuracy and generalization, while also decreasing computation time in plant disease detection systems.
Comparative Analysis of Existing Techniques
A comparative analysis of the existing plant disease detection approaches provides insight into model strengths, limitations, and performance trade-offs. Table 3 summarizes the accuracy, core model characteristics, and key limitations of classical, quantum, and hybrid DL frameworks.
Table 3: Comparative Analysis of Existing Plant Disease Detection Approaches
| Approach | Core Model | Enhancement | Accuracy (%) | Key Limitation |
| CNN6 | Convolutional layers | None | 88–90 | Overfitting, slow training |
| DenseNet8 | Dense connectivity | Feature reuse | 91–93 | High memory use |
| CapsNet + Attention11 | Capsule routing + attention | Spatial feature focus | 92–94 | Complex routing mechanism |
| QNN12 | Quantum variational circuits | Quantum encoding | 93–95 | Hardware constraints |
| Hybrid (DenseNet + Attention + CapsNet + QNN + ECP-HSOA)22 | Hybrid feature fusion | Quantum-inspired optimization | 95+ | Implementation complexity |
The analysis focuses on classical CNNs offering a solid starting point in feature extraction, but when it comes to real-world applications, the generalization problem becomes even more apparent and pronounced. Density improves feature reuse and gradient flow but has high memory requirements. Capsule Networks combined with attention networks offer interpretability and improved spatial feature representation but at a greater computational cost. Though QNNs provide high-dimensional embeddings and achieve faster convergence, they pass the convergence on to quantum-hardware-restricted dimensionality. Through the hybrid models described, which integrate classical and quantum models with ECP-HSOA, the described models achieve the best accuracy and efficiency, but they require more work to implement.
Challenges and Research Gaps
Despite notable advancements, several challenges persist in applying quantum-enhanced DL for plant disease detection. Table 4 summarizes the major obstacles and research gaps identified in the literature.
Table 4: Challenges and Research Gaps in Quantum-Enhanced DL for Plant Disease Detection
| Challenge | Description | Research Gap |
| Quantum hardware limitations | Limited qubits and high noise levels restrict large-scale applications | Development of robust, scalable quantum processors |
| Data encoding complexity | Efficient conversion of high-dimensional leaf images into quantum states is non-trivial | Research into efficient quantum feature mapping |
| Algorithmic integration | Hybrid classical–quantum models require careful tuning of learning rates, loss functions, and optimization strategies | Standardized frameworks for stable hybrid model integration |
| Explainability | Lack of interpretability for quantum-enhanced features | Development of explainable quantum AI techniques for agriculture |
| Computational efficiency | Classical DL models are computationally intensive | Lightweight QNN architectures for edge deployment |
Addressing these challenges is essential to enable practical deployment in precision agriculture. Future research should focus on lightweight hybrid architectures, explainable quantum layers, and edge-compatible quantum simulators to facilitate real-world applications.
Future Directions
Emerging trends suggest several promising directions for quantum-enhanced DL in agriculture:
Quantum Edge Computing – Implementing quantum-enhanced inference on edge devices allows on-field plant disease detection, reducing latency and reliance on cloud processing.28
Explainable Quantum AI (XQAI) – Combining interpretability techniques such as SHAP or LIME with quantum feature visualization can increase trust and adoption among farmers and agronomists.
Quantum Transfer Learning – Utilizing pretrained classical models to initialize hybrid quantum architectures reduces computational overhead and accelerates convergence.29
Federated Quantum Learning – Distributed, privacy-preserving training across multiple agricultural datasets can enable collaborative intelligence without compromising sensitive data.
These directions emphasize the integration of scalability, efficiency, and interpretability, ensuring that quantum-enhanced DL systems are practical and deployable in diverse agricultural settings.
Discussion
The review highlights that deep learning-based plant disease detection models have achieved significant progress, particularly in improving classification accuracy and robustness across diverse crop species. Architectures such as CNNs, Dense Nets, and Capsule Attention Networks have demonstrated strong feature extraction capabilities and resilience to image distortions. However, the practical scalability and real-time deployment of these models remain constrained by their computational complexity, memory requirements, and sensitivity to environmental variations such as lighting, leaf angle, and background noise. As agriculture increasingly transitions toward data-driven, automated systems, reliance on large-scale GPUs or cloud-based inference becomes impractical for rural or decentralized farm environments.
Quantum-enhanced deep learning frameworks provide a promising direction for addressing these limitations. The ability of quantum neural networks (QNNs) to encode data into high-dimensional Hilbert spaces and exploit quantum principles such as superposition and entanglement may enable more expressive learning capacity with fewer parameters compared to classical networks. Hybrid models that integrate quantum layers with classical feature extractors show potential for improved convergence stability and reduction in training time, particularly when combined with quantum-inspired optimization techniques such as ECP-HSOA. These hybrid architectures may be especially valuable for tasks requiring discrimination between visually similar disease patterns or early-stage infections where classical models struggle to generalize.
However, several open challenges persist. The deployment of quantum-enhanced frameworks in real agricultural settings is limited by quantum hardware immature, noise instability, and the scarcity of qubit resources. Data encoding remains a core bottleneck, as converting high-resolution leaf imagery into quantum states introduces additional computational overhead. Moreover, current models lack interpretability: while quantum layers can boost performance, their decision-making process is often opaque, making it difficult for agronomists and plant pathologists to trust or validate model outputs. This emphasizes the need for explainable quantum AI (XQAI) techniques to bridge the interpretability gap and ensure domain-level knowledge integration.
Another notable research gap is the absence of standardized evaluation benchmarks across real-field datasets. Most reported results are derived from controlled laboratory image collections, limiting generalizability. Future research must prioritize field-based datasets, multimodal sensing (combining hyperspectral, thermal, and drone imaging), and longitudinal monitoring of disease progression. Furthermore, resource-efficient edge-deployable hybrid frameworks should be developed to ensure real-time inference directly on low-power smart farming devices, minimizing reliance on high-end computing infrastructure.
In summary, while quantum-enhanced deep learning frameworks hold considerable promise for transforming plant disease detection, the path toward operational-scale adoption requires addressing challenges related to hardware feasibility, data representation, interpretability, and system integration. Advancing interdisciplinary collaboration among quantum computing researchers, agricultural scientists, and computational biologists will be essential for developing sustainable, reliable, and scalable intelligent agriculture systems.
Conclusion
The survey showcases the groundbreaking capabilities of quantum-enhanced deep learning in detecting plant diseases. Integrating QNN layers with classical architectures like DenseNet and Capsule Networks and optimizing those using ECP-HSOA yields improvements in accuracy, convergence speed, and computational efficiency
Meanwhile, the hybrid frameworks have exceeded 95% accuracy in classification tasks, surpassing traditional CNNs by 5% and trimming the training period by over 30%. This progress facilitates accurate leaf segmentation, comprehensive disease classification, and flexibility in changing environmental conditions.
Currently, research ought to concentrate on the creation of lightweight, interpretable, and quantum-enhanced deep learning models that are suitable for deployment on the edge. Such frameworks have the potential to provide intelligent, sustainable, and scalable systems for agricultural monitoring and will form the basis of the next generation of smart farming.
Acknowledgement
The authors would like to express their sincere gratitude to the Department of Computer Science & Technology, Sri Krishnadevaraya University, Anantapuramu, Andhra Pradesh, India, for providing continuous support, encouragement, and the necessary research facilities to conduct this study. The constructive feedback and guidance from colleagues and mentors greatly contributed to the completion of this work.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The authors do not have any conflict of interest.
Data Availability Statement
No new data were generated or analyzed in this study. Data sharing is not applicable to this article as it is a survey-based work that synthesizes information from previously published studies.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Permission to Reproduce Material from Other Sources
Not Applicable.
Author Contributions
Usikela Naresh: Conceptualized the study, Conducted the literature survey, and Prepared the initial manuscript draft.
Thota Bhaskar Reddy: Supervised the research, Provided critical revisions, and Contributed to the final review and editing of the manuscript.
References
- Kulkarni K. P., Vennapusa A. R., Pandian B. A., Deshmukh R. Genetic advancements for improving plant tolerance to biotic and abiotic stresses. Front Genet. 2024;15:1426680.
CrossRef - Mehedi M. H. K., Nawer N., Ahmed S., Khan M. S. I., Hasib K. M., Mridha M., Alam M. G. R., Nguyen T. T. PLD-Det: Plant leaf disease detection in real time using an end-to-end neural network approach based on improved YOLOv7. Neural Comput Appl. 2024;36:21885–21898.
CrossRef - Nyawose T., Maswanganyi R. C., Khumalo P. A review on the detection of plant disease using machine learning and deep learning approaches. J Imaging. 2025;11(10):326. https://doi.org/10.3390/jimaging11100326
CrossRef - Tatana M. M., Tsoeu M. S., Maswanganyi R. C. Low-light image and video enhancement for more robust computer vision tasks: A review. J Imaging. 2025;11:125.
CrossRef - Bhujade V. G., Sambhe V., Banerjee B. Digital image noise removal towards soybean and cotton plant disease using image processing filters. Expert Syst Appl. 2024;246:123031.
CrossRef - Li Z., Fan X., Wang Q., Xu W. Quantum-enhanced algorithms for image classification: a case study on agricultural datasets. J Quantum Inf Sci. 2022;12(2):55–70.
- Mohanty S. P., Hughes D. P., Salathé M. Using deep learning for image-based plant disease detection. Front Plant Sci. 2016;7:1419. doi:10.3389/fpls.2016.01419
CrossRef - Hadiq H., Solehatin S., Djuniharto D., Muslim M. A., Salahudin S. N. Comparison of the suitability of the Otsu method thresholding and multilevel thresholding for flower image segmentation. J Soft Comput Explor. 2023;4:242–249.
CrossRef - Sarkar C., Gupta D., Gupta U., Hazarika B. B. Leaf disease detection using machine learning and deep learning: Review and challenges. Appl Soft Comput. 2023;145:110534.
CrossRef - Preskill J. Quantum computing in the NISQ era and beyond. Quantum. 2018;2:79. doi:10.22331/q-2018-08-06-79
CrossRef - Schuld M., Killoran N. Quantum machine learning in feature Hilbert spaces. Phys Rev Lett. 2019;122(4):040504.
CrossRef - Punitha K., Shrivastava G. Analytical study of deep learning architectures for classification of plant diseases. J Algebr Stat. 2022;13:907–914.
- Amarjeeth S., Jaisree H., Aishwarya D., Jayasree J. S. Plant disease detection and diagnosis using deep learning. In: Proc Int Conf for Advancement in Technology (ICONAT). Goa, India; January 21–22, 2022:1–6.
CrossRef - Vishakha K., Munot M. Deep learning models for tomato plant disease detection. In: Advanced Machine Intelligence and Signal Processing. Singapore: Springer; 2022:679–686.
CrossRef - Khalil K., Khan R. U., Albattah W., Qamar A. M. End-to-end semantic leaf segmentation framework for plant disease classification. Complexity. 2022;55:1076–2787.
CrossRef - Wadadare S. S., Fadewar H. S. Deep learning convolution neural network for tomato leaves disease detection by inception. In: Applied Computational Technologies: Proc ICCET. Singapore: Springer; 2022:208–220.
CrossRef - Chen H. C., Widodo A. M., Wisnujati A., Rahaman M., Lin J. C. W., Chen L., Weng C. E. AlexNet convolutional neural network for disease detection and classification of tomato leaf. Electronics. 2022;11:951.
CrossRef - Khan M. A., Alqahtani A., Khan A., Alsubai S., Binbusayyis A., Ch M. M. I., Yong H. S., Cha J. Cucumber leaf diseases recognition using multi-level deep entropy–ELM feature selection. Appl Sci. 2022;12:593.
CrossRef - Xiting Y., Plett D., Liu H. Detecting crown rot disease in wheat in controlled environment conditions using digital color imaging and machine learning. AgriEngineering. 2022;4:141–155.
CrossRef - Kaur P., Harnal S., Tiwari R., Upadhyay S., Bhatia S., Mashat A., Alabdali A. M. Recognition of leaf disease using hybrid convolutional neural networks with feature reduction. Sensors. 2022;22:575.
CrossRef - Almadhor A., Rauf H. T., Lali M. I. U., Damaševičius R., Alouffi B., Alharbi A. AI-driven framework for recognition of guava plant diseases through machine learning from DSLR camera–based high-resolution imagery. Sensors. 2021;21:3830.
CrossRef - Applalanaidu M. V., Kumaravelan G. A review of machine learning approaches in plant leaf disease detection and classification. In: Proc Third Int Conf on Intelligent Communication Technologies and Virtual Mobile Networks (ICICV). Tirunelveli, India; February 4–6, 2021:716–724.
CrossRef - Satheesh S. B. P., Naidu P. R. S., Neelima U. Prediction of guava plant diseases using deep learning. In: Proc 3rd Int Conf on Communications and Cyber Physical Engineering (ICCCE). Hyderabad, India; April 9–10, 2021:1495–1505.
CrossRef - Murat T., Arslan R. S. RT-Droid: A novel approach for real-time Android application analysis with transfer learning–based CNN models. J Real-Time Image Process. 2023;20:55.
CrossRef - Al-Gaashani M. S., Shang F., Muthanna M. S., Khayyat M., Abd El-Latif A. A. Tomato leaf disease classification by exploiting transfer learning and feature concatenation. IET Image Process. 2022;16(3):913–925.
CrossRef - El Akhal H., Yahya A. B., Moussa N., El Alaoui A. E. B. Image-based olive leaf disease classification using a deep hybrid model. Ecol Inform. 2023;77:102276.
CrossRef - Ferentinos K. P. Deep learning models for plant disease detection and diagnosis. Comput Electron Agric. 2018;145:311–318.
CrossRef - Ghodekar A., Kumar A. LightLeafNet: lightweight and efficient NASNetMobile architecture for tomato leaf disease classification. SSRN. 2023. Available at: https://ssrn.com/abstract=4570347
CrossRef - Hamim S. A., Jony A. I. Enhancing brain tumor MRI segmentation accuracy and efficiency with optimized U-Net architecture. Malays J Sci Adv Technol. 2024;4(3):197–202.
CrossRef

