Introduction
Phosphorus is a needed supplement for the development and progress of plant life and progress, yet its attainability in the soil is often slight due to its appearance in indissoluble forms. This scarcity of bioavailable phosphorus poses a significant challenge to plant nutrition, particularly in agricultural soils. Phosphate solubilizing bacteria (PSBs) act as a fundamental factor in addressing this challenge by converting insoluble phosphorus compounds into soluble forms that plants can effectively absorb. Agroecosystems that improve soil fertility and crop productivity, especially in nutrient-deficient regions, have gained critical importance, as highlighted by numerous studies.1 Microbial cultures of PGPRs are advantageous soil microbes that inflate plant growth through various mechanisms, one of which is the solubilization of phosphate. These bacteria secrete organic acids, phosphatases, and other compounds (figure 1), that help release phosphorus into a form plant can utilize.2 Besides improving phosphorus availability, PSBs contribute to overall soil health and fertility. By solubilizing phosphate, they enhance nutrient uptake and foster beneficial interactions with plant roots. This symbiotic relationship is crucial for plant growth and progress, particularly under phosphorus-limited provisions. Some PSBs also produce some phytohormones such as gibberellic acid, auxins, and cytokinins, further promoting root development and plant growth.3
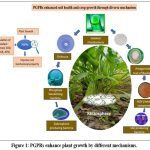 |
Figure 1: PGPRs enhance plant growth by different mechanisms. |
The role of PGPR in phosphate solubilization is especially significant in regions with phosphorus-deficient soils, where traditional fertilization practices may not be sustainable or environmentally friendly. By leveraging the natural abilities of PSBs to enhance nutrient availability, agronomists can abate the need for chemical compost, and assist in more sustainable farming. Recent studies emphasize the contribution of PSBs such as Bacillus sp., Pseudomonas sp., and Burkholderia sp., in ameliorating soil fertility and yield of crops across diverse agricultural systems.4
Iron (Fe) is an essential micronutrient for plants, playing a pivotal role in various physiological processes such as chlorophyll synthesis, respiration, and photosynthesis.5 Despite its abundance in the Earth’s crust, iron’s bioavailability to plants is often limited due to its tendency to form insoluble hydroxides and oxides under aerobic soil conditions, leading to iron deficiency in plants6 to overcome this limitation, certain soil microorganisms, notably plant growth-promoting rhizobacteria (PGPR), produce siderophores low molecular weight, high-affinity iron-chelating compounds that solubilize and sequester iron from the soil, making it accessible to plants.7
Siderophores not only facilitate iron uptake but also play a significant role in suppressing soil-borne pathogens by outcompeting them for iron, thereby acting as biocontrol agents. The production of siderophores by PGPR such as Pseudomonas, Bacillus, and Azotobacter species has been extensively documented, with studies highlighting their efficacy in enhancing plant growth and resilience under various abiotic stresses, including salinity and drought.8
The purpose of this study was to isolate and screen the phosphate-solubilizing and siderophore-producing PGPRs from the rhizospheric region of Solanum lycopersicum and to evaluate the impact of these bacteria on phosphorus release and availability in the soil. By understanding and utilizing the phosphate-resolubilizing capabilities of PGPR, this research aims to enhance plant growth and support sustainable agricultural practices.
While numerous studies have reported the utility of PGPRs in enhancing nutrient availability and plant growth, most research has centered on well-studied genera such as Pseudomonas, Bacillus, and Azospirillum.9 Comparatively, the potential of Burkholderia spp. as multifunctional PGPRs remains underexplored, particularly in the context of rhizosphere soils from tomato cultivation systems in underrepresented agroecological zones. This study addresses this research gap by isolating and characterizing a novel strain of Burkholderia dolosa (RBS11) from the tomato rhizosphere, with a dual focus on its phosphate-solubilizing and siderophore-producing capabilities. The isolate was subjected to a comprehensive in vitro screening to assess key PGPR traits prior to molecular identification via 16S rRNA sequencing and phylogenetic analysis. Importantly, this study does not extend to in planta or field evaluation of the isolate’s bioefficacy. Hence, the scope is limited to laboratory-based functional profiling, providing a basis for future greenhouse and field trials.
This research contributes to the expanding knowledge on lesser-known PGPR genera and their biotechnological potential in sustainable agriculture, particularly in phosphorus-deficient regions. The novelty lies in the dual-functional profiling of B. dolosa (RBS11), a strain not widely reported for such traits, and its contextual relevance to tomato rhizosphere microbiomes.
Material and Methods
Sample collection
The soil sample adhered to the roots of tomato rhizosphere were collected from the field. The plants were collected from 2 different locations of Uttarakhand viz., farmer field of Roorkee and Bahadrabad. The plants were carefully uprooted from soil and soil adhering to the root was put in poly bags for isolation of rhizobacteria.
Isolation of rhizobacteria
In addition to isolating bacteria using a selective medium (King’s B medium), the medium typically consists of 20g of protease peptone, which serve as a rich nitrogen source, 1.5g of dipotassium hydrogen phosphate (K2HPO4) to maintain buffering capacity, 1.5g of magnesium sulfate heptahydrate (MgSO47H2O) providing essential magnesium ions, and 10ml of glycerol as a carbon and energy source. The components are dissolved in 1 liter of distilled water and pH is 7.2 before autoclaving.
One gram of the collected rhizospheric soil sample is suspended in 9ml of sterile distilled water and serially diluted up to 10-6. Aliquots from appropriate dilutions are spread on selective or general- purpose media such as King’s B medium and Pikovskaya’s agar and were analyzed for PSB (Phosphate solubilizing bacteria), which plays a crucial role in enhancing plant growth by converting insoluble phosphate forms into bioavailable forms. The soil samples were inoculated on Pikovskaya’s agar medium, a selective medium used to detect PSB after incubation of 48 hours at 28o C clear halo zones sounding the microbial colonies indicated phosphate solubilization activity. These isolated bacterial colonies were then purified by repeated streaking on Pikovskaya’s medium until pure cultures were obtained. The physiological characteristics, such as bacterial colony shape, bacterial size, color, and the extent of halo formation, were carefully observed to select different PSB strains for further characterization.10
Qualitative assay
By streaking the bacterial cultures across the Pikovskaya medium surface containing the following components per liter: glucose (10 g), tricalcium phosphate (5 g), ammonium sulfate (0.5 g), potassium chloride (0.2 g), magnesium sulfate (0.1 g), yeast extract (0.5 g), manganese sulfate (0.0001 g), ferrous sulfate (0.0001 g), and agar (15 g); pH adjusted to 7.0 before autoclaving. Plates were incubated at 28°C for 48 hours. appearance of a clearing zone surrounding the bacterial colony was utilized as a sign of positive phosphate solubilization (Figure 2).11
Quantitative assay
The quantitative assessment of phosphate solubilization in a medium containing an insoluble form of phosphate was examined in Pikovskaya’s broth following incubation at 28±1°C for 72 hours. The pH of the bacterial culture broth was resolute, and the culture was centrifuged at 9,000 rpm for 15 minutes at 4°C. The amount of solubilized tricalcium phosphate in the supernatant was resolute using an alter method based on Pikovskaya (1948), with updated procedures for accuracy.12 The production profile was observed by estimating the amount of solubilized tricalcium phosphate over time.
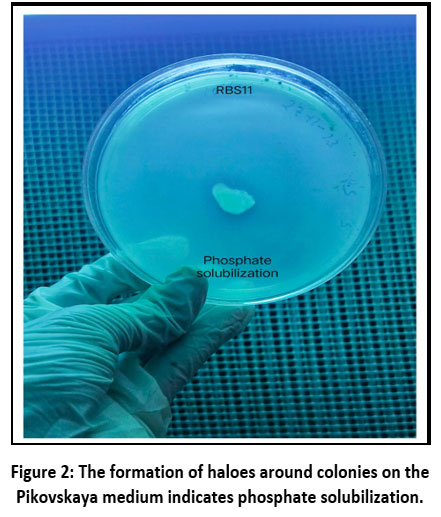 |
Figure 2: The formation of haloes around colonies on the Pikovskaya medium indicates phosphate solubilization. |
Siderophore estimation assay
Qualitative assay
Siderophore production was assessed on Chrome Azurol S (CAS) agar (Schwyn & Neiland, 1987). CAS agar was prepared freshly by combining the CAS dye solution, iron (III)-hexadecyltrimethylammonium bromide (Fe-HDTMA) complex, and minimal salt medium (MM9 base). All glassware was pre-treated by soaking in 6 M HCl overnight to eliminate iron contamination and rinsed thoroughly with distilled water. Bacterial isolates were spot inoculated on CAS agar and incubated at 28°C for 48–72 hours. The development of orange halos around the colonies indicated positive siderophore production (Figure 3).
consisting of 0.5 mL of CAS reagent and 1 mL of uninoculated broth, was prepared for comparison. The siderophore concentration in each sample was determined using the formula:
% Siderophore Units = (Ar-As)/Ar
Where Ar is the absorbance at 630 nm of the reference solution (CAS assay solution + uninoculated media) and As is the absorbance at 630nm of the sample (CAS assay solution + supernatant)13
Morphological and biochemical characterization
The strains were assessed for their physiological features according to the guidelines outlined in Bergey’s Manual of Determinative Bacteriology. This change encompassed examining cell morphology, configuration, Gram staining attributes, and various biochemical properties, including hydrogen cyanide (HCN) production, nitrogen fixation, zinc dissolution, ammonia prolongation, and auxin synthesis. Additionally, the analysis involved measuring enzymatic activities, particularly the prolongation of chitinase and protease.14
 |
Figure 3: The presence of orange halos surrounding colonies on blue agar indicates the secretion of siderophores. |
Molecular characterization
For molecular identification, one promising rhizobacteria was selected for further identification. The genomic DNA of the selected isolate (RBS11) was prepared individually as described by.15 MEGA 11 software was used to make out the phylogeny. The accession numbers were obtained by submitting the gene sequence of 16S rRNA to NCBI GenBank Database.
In-vitro screening for Antagonistic traits
The antagonistic activity of phosphate-solubilizing and siderophore-producing bacteria was evaluated against Fusarium oxysporum, the pathogen responsible for Fusarium wilt in Solanaceous crops, using the dual culture method. In this experiment, the antagonistic bacterial strains were streaked onto potato dextrose agar (PDA) plates, ensuring a 2 cm distance from the pre-placed fungal mycelium (3 mm in diameter). They were then put into the BOD incubator to incubate it at 28°C for 7 days. After incubation, the radial growth of the fungal colony was measured. Fungal growth inhibition was deliberate by the formula:
% of inhibition = (R-r) / R ×100
(r) represents the radius of the pathogenic fungi in the presence of the isolated bacteria colony, and R denotes the maximum radius of the pathogenic fungi without bacterial interference.
Statistical analysis
Data analysis was done using analysis of variance (ANOVA) with the Statistics 8.1 software package. The importance of conduct toward means became evaluated and differentiated the usage of the least sizeable difference (LSD) test.16
Results
Isolation and screening
A total of fifty unique bacterial isolates were collected after enriching and isolating samples from the rhizospheric soil of lycopersicum cultivation in the Haridwar region. These isolates designated RBS-1 to RBS-50, were characterized individually for their abilities to solubilize inorganic phosphate of soil and fabricate siderophores.
Screening for phosphate production
Qualitative Assay
Phosphate solubilization is a crucial characteristic of rhizospheric bacteria (PGPRs) and it has gained importance in recent years for its potential applications beyond agriculture. In this study, we examined 50 bacterial strains obtained from the rhizospheric region of Solanum lycopersicum for their ability to solubilize phosphate using Pikovskaya agar plates. A zone around the rhizobacterial colonies indicated the transformation of insoluble phosphate into a soluble form. Out of the 50 isolates tested, 10 were identified as positive for phosphate solubilization. The isolates RBS-4, RBS-7, RBS-11, RBS-13, RBS-14, RBS-20, RBS-23, RBS-38, RBS-40, and RBS-50 demonstrated the highest phosphate solubilizing capabilities, as evidenced by the size of the halo zones on the Pikovskaya agar plates (see Table 1, figure 4).
Quantitative assay
Since Pikovskaya medium exhibited a greater pH decrease (Figure 6), compared to the other media, it was utilized for the quantitative assessment of phosphate solubilization by the isolates RBS4, RBS7, RBS11, RBS13, RBS14, RBS20, RBS23, RBS38, RBS40, and RBS50 using the Pikovskaya method. In this process, a maximum of 92 mg/L of phosphate was observed to be solubilized from tricalcium phosphate by RBS11 (Table 1).
Table 1: Qualitatively and quantitatively assessment of phosphate solubilizing by bacterial strains.
| S. No | Isolated code | Qualitative | pH | Quantitative (Phosphate solubilization) |
S. No | Isolates code | Qualitative | pH | Quantitative (Phosphate solubilization) |
| 1 | RBS-4 | + | 4.2+/-0.1 | 12+/-2.23 | 6 | RBS-20 | + | 4.1+/-0.1 | 18+/-3.21 |
| 2 | RBS-7 | + | 4.5+/-0.1 | 83+/-2.98 | 7 | RBS-23 | + | 3.5+/-0.1 | 83+/-3.23 |
| 3 | RBS-11 | + | 3+/-0.1 | 92+/-3.21 | 8 | RBS-38 | + | 4.5+/-0.1 | 32+/-4.31 |
| 4 | RBS-13 | + | 4+/-0.1 | 25+/-4.81 | 9 | RBS-40 | + | 4.3+/-0.1 | 25+/-4.81 |
| 5 | RBS-14 | + | 4+/-0.1 | 32+/-2.14 | 10 | RBS-50 | + | 4+/-0.1 | 13+/-4.25 |
Abbreviation: + = Positive, pH = Potential pf Hydrogen, ± = Standard deviation from mean value.
Assessment of siderophore synthesis.
Qualitative assay
Production of siderophore is an essential characteristic of PGPRs and has received considerable interest in recent decades due to its applications beyond agricultural contexts. Utilizing CAS agar plates, this work evaluated all 50 rhizobacterial isolates obtained from the rhizosphere of Solanum lycopersicum for their ability to produce siderophores. The proximity of an orange clear zone around the bacterial colonies recommended siderophore synthesis. Out of the 50 isolates, only 10 showed positive results for siderophore production. Isolates RBS-4, RBS-7, RBS-11, and RBS-50 demonstrated the highest levels of siderophore production in the CAS agar medium (Table 2). Siderophore production was estimated based on the diameter of the halo formed on the CAS agar plates.
Table 2: Qualitatively and quantitatively assessment of siderophore synthesis from rhizobacterial isolates.
| S. No | Isolates code | Qualitative | Quantitative (%SU) |
S. No | Isolates code | Qualitative | Quantitative (%SU) |
| 1 | RBS-4 | +++ | 60 | 6 | RBS-20 | ++ | 41 |
| 2 | RBS-7 | +++ | 62 | 7 | RBS-23 | ++ | 60 |
| 3 | RBS-11 | +++ | 70 | 8 | RBS-38 | + | 32 |
| 4 | RBS-13 | ++ | 42 | 9 | RBS-40 | + | 35 |
| 5 | RBS-14 | ++ | 48 | 10 | RBS-50 | + | 38 |
Abbreviation: +++ = Strong siderophore production, ++ = Moderate siderophore production, + = Week siderophore production.
Quantitative assay
The quantitative assessment of siderophore synthesis was conducted using liquid culture media along with CAS reagent, with results expressed in siderophore units (SU%). The analysis indicated that the ten bacterial isolates generated siderophore units ranging from 32% to 70% after 48 hours of incubation (see Table 2). The highest amount of siderophore production, at 70%, was observed for isolate RBS-11, which was statistically significant compared to the other isolates. Isolate RBS-7 followed, yielding 62% SU, and was statistically similar to RBS-11. The other isolates showed lower levels of iron carrier origination but were crucially different from the control group. Among these, RBS-38 and RBS-40 exhibited the least synthesis of siderophores (Figure 5).
Morphological and biochemical characteristics of selected rhizobacterial isolates.
Based on the features identified through Gram staining, 10 isolates were classified as gram-positive, while 8 were determined to be gram-negative. Among the siderophore-producing bacterial isolates, two distinct morphological types were noted: cocci and bacilli (see Table 3). Additional analysis of the 10 siderophore-producing isolates shows their implicit for various plant growth-promoting activities. All isolates tested positive to produce phytohormone (IAA). Moreover, nine rhizobacterial isolates demonstrated positive results for zinc solubilization, and on the Pikovskaya medium, all 10 isolates created clear halo zones (Table 3).
Table 3: Morphological and biochemical Screening of Siderophore and phosphate solubilizing strains for plant growth-promoting activities.
| S. No | Isolates naming | Shape of isolates | Gram nature of isolates | PSBs | Auxin | Zinc | Chitinase | Protease |
| 1 | RBS-4 | Bacilli | – | + | + | + | + | + |
| 2 | RBS-7 | Bacilli | + | + | + | + | + | + |
| 3 | RBS-11 | Bacilli | – | + | + | + | + | + |
| 4 | RBS-13 | Cocci | – | + | + | + | + | + |
| 5 | RBS-14 | Cocci | + | + | + | + | + | + |
| 6 | RBS-20 | Bacilli | + | + | + | + | + | + |
| 7 | RBS-23 | Bacilli | + | + | + | + | + | + |
| 8 | RBS-38 | Bacilli | – | + | + | + | + | + |
| 9 | RBS-40 | Bacilli | – | + | + | + | + | + |
| 10 | RBS-50 | Cocci | + | + | + | + | + | + |
Abbreviation; + = Positive, – = Negative.
 |
Figure 4: Quantitative assessment of isolated Phosphate solubilizing rhizobacteria. |
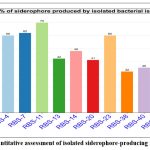 |
Figure 5. Quantitative assessment of isolated siderophore-producing rhizobacteria. |
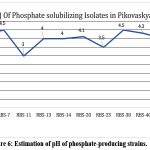 |
Figure 6: Estimation of pH of phosphate-producing strains. |
Additionally, all phosphate- and siderophore-producing isolates demonstrated significant chitinase production, an important characteristic of PGPR. Among the isolates, three tested positives for hydrogen cyanide (HCN) production. Regarding hydrolytic enzyme activity, only three out of the ten isolates showed positive results for protease production.
In-vitro Screening for Antagonistic Properties
Among the 10 rhizobacterial isolates tested, two isolates demonstrated inhibitory activity against Fusarium oxysporum. These were RBS-7, which exhibited 35% inhibition, and RBS-11, which showed the highest inhibition at 46%. The most significant inhibitory effect was observed with RBS-11, forming a 3 mm inhibition zone (Figure 7). Given its superior inhibition potential, RBS-11 can be considered a promising candidate for biocontrol applications against Fusarium oxysporum.17
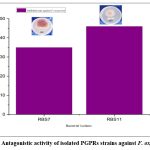 |
Figure 7: Antagonistic activity of isolated PGPRs strains against F. oxysporum. |
Molecular characterization
The effective bacterial isolates (RBS11) that solubilized insoluble form of phosphate in soil and synthesized siderophores were selected and were identified as Burkholderia dolosa strains having 97 % similarity with the reported gene sequence (Table 4). The resulting phylogenetic tree depicts the evolutionary relationship of the selected isolate with closely related, validly named species (Figure 8). The nucleotide accession numbers for strain RBS11 are PQ059698.
Table 4: Identification of isolated strains based on 16S rRNA gene sequence analysis
| Bacterial strain | Genus | Closely related validly published species |
Taxon | NCBI Number (found after sequence submitted in NCBI database |
| RBS-11 | Burkholderia | Burkholderia dolosa | b-proteobacteria | PQ059698 |
 |
Figure 8: Neighbor-joining phylogenetic tree based on 16S rRNA gene sequences, showing the phylogenetic position of the selected strain RBS-11. |
Discussion
The rhizosphere is widely recognized as a hotspot of microbial activity where plant growth-promoting rhizobacteria (PGPR) play a pivotal role in enhancing crop productivity, particularly under nutrient-limiting conditions.18 Phosphorus and iron, essential for plant metabolism, often remain unavailable due to their insoluble forms in soil, necessitating microbial intervention for mobilization.19 PGPR such as Pseudomonas, Bacillus, and Burkholderia have demonstrated significant potential in solubilizing inorganic phosphate through the production of organic acids and phosphatases20 while simultaneously producing siderophores that chelate ferric iron and enhance its bioavailability.21 The dual functionality of these bacteria has been validated through qualitative and quantitative assays such as Pikovskaya’s medium for phosphate solubilization and CAS agar/liquid assays for siderophore detection.22 In the present study, isolates RBS-11 and RBS-7 from the rhizosphere of tomato plants cultivated in iron- and phosphate-deficient soils of Haridwar showed promising results, with RBS-11 producing 70% siderophore units and solubilizing 92 mg/L of phosphate, indicating its potential as an efficient biofertilizer.23 Similar findings have been reported for tomato rhizobacteria under saline and alkaline soil conditions, suggesting that such PGPR improve nutrient uptake, root development, and resistance to stress.24,25 The application of PGPR-based consortia has also been shown to enhance the availability of other micronutrients while promoting soil fertility and suppressing phytopathogens. Modern molecular and metagenomic tools have enabled the identification of key functional genes associated with phosphate solubilization (e.g., pqqE, gcd) and siderophore biosynthesis (e.g., iucA, entD), offering deeper insight into rhizobacterial functions.26,27 Recent advances have emphasized the need for integrating such beneficial strains into sustainable agricultural practices to reduce chemical fertilizer dependency and maintain ecological balance.28,29 Overall, the dual action of these PGPR offers a promising biological route to alleviate phosphate and iron deficiencies in tomato cultivation while enhancing plant growth sustainably.30 Building upon the promising results of this study, future research should aim to conduct whole-genome sequencing and comparative genomics of potent isolates such as RBS-11 and RBS-7 to identify31,32 key genes involved in phosphate solubilization and siderophore biosynthesis.33 This will provide deeper insights into the molecular mechanisms underlying their plant growth-promoting capabilities.34,35 Additionally, greenhouse and open-field trials under diverse agro-climatic conditions should be undertaken to validate the efficacy and consistency of these strains in enhancing tomato productivity.36 It is also essential to explore the potential synergistic effects of microbial consortia, combining phosphate-solubilizing, nitrogen-fixing, and biocontrol PGPR for a more holistic and sustainable approach to crop nutrition.37 Moreover, studies focusing on formulation development and shelf-life stabilization of these bacterial isolates are crucial for their commercialization as biofertilizers.38 Integration of metagenomics and transcriptomics will further help in understanding the interaction dynamics between introduced strains and native microbial communities in the rhizosphere. Lastly, the environmental and economic impacts of long-term application of these bioinoculants should be assessed to promote their adoption in organic and sustainable farming systems.39
Conclusion
This study identified and characterized phosphate-solubilizing and siderophore producing bacteria from the rhizosphere of tomato plants in solubilization of phosphate and iron- deficient soils. RBS-11 emerged as the most promising candidate among the isolates due to its high siderophore production and phosphate solubilization ability. These revelations suggest that these bacteria could play an essential role in ameliorating the nutrient uptake efficiency of tomato plants, potentially enhancing crop yields in nutrient-poor soils. Further studies focusing on these isolates’ field application and their influence on plant growth soil health and fertility are needed to confirm their potential as biofertilizers in sustainable agriculture.
Acknowledgement
The author would like to thank GuruKula Kangri (Deemed to be) University, Haridwar for granting the Ph.D. research work. The Department of Botany and Microbiology is highly appreciated for allowing the laboratory work.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The authors do not have any conflict of interest.
Data Availability Statement
The manuscript incorporates all datasets produced or examined throughout this research study.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Author Contributions
Sadhana Giri; Conceptualization, Methodology, writing – Original Draft, Data Collection, Analysis, Writing – Review & Editing.
Verinder Virk; Visualization, Supervision, Project Administration.
References
- Banerjee S, Sarkar S, Mitra S. Biofertilizer potential of multifunctional PGPR for sustainable tomato cultivation. Curr Microbiol. 2024;81(1):45-56.
- Bano A, Asad SA, Rizvi ZF. Characterization of PGPR from tomato rhizosphere for phosphate solubilization and siderophore production. Microbiol Res. 2024; 267:127311.
- Choudhary D, Sharma A, Singh R. Bioformulations of rhizobacteria for sustainable crop production. J Soil Sci Plant Nutr. 2025;25(2):200-214.
- Ehinmitan E, Siamalube B, Mamati E, Ngumi V, Juma P, Losenge T. Evaluating growth-promotion and drought tolerance properties of endophytic Methylobacterium spp. from semi-arid Kenya soil. Scope. 2024;14(3):924-940.
- Fernandes G, Leite M, Souza D. Siderophore-mediated iron acquisition by PGPR under iron-deficient soil conditions. Biotechnol Lett. 2024;46(3):345-357.
- Ferreira FN, Lima GSD, Gheyi HR, Sá FVD, Dias AS, Pinheiro FW. Photosynthetic efficiency and production of Annona squamosa L. under salt stress and fertilization with NPK. Rev Bras Eng Agric Ambient. 2021;25(7):446-452.
CrossRef - Ghosh A, Dey N, Bhattacharyya P. Role of phosphate solubilizing bacteria in sustainable agriculture: An overview. Sustain Agric Res. 2022;10(4):64-72.
- Gupta R, Jain V, Tiwari S. Phosphate-solubilizing bacteria from tomato rhizosphere: Soil nutrient enhancement under stress. Plant Soil. 2024;489:255-269.
- Jha BK, Bharti A, Kumar A. Phosphate-solubilizing and siderophore-producing bacteria from tomato: A dual approach. World J Microbiol Biotechnol. 2023;39(7):118.
- Joshi D, Kumar V, Singh M. Rhizobacterial interventions for sustainable tomato farming. Agric Microbiol. 2024;42(1):1-12.
- Kaur T, Verma M, Reddy MS. Evaluation of phosphate solubilizing bacteria for phosphate solubilization efficiency and plant growth-promoting traits. Environ Sustain. 2022;5(3):195-205.
- Kour D, Rana KL, Yadav AN. PGPR bioformulations and the future of agro-biotechnology. Agric Microbiol. 2023;41(3):201-215.
- Kumar A, Maurya BR, Raghuwanshi R. Characterization of siderophore-producing rhizobacteria for enhancing plant growth and nutrient uptake under iron-limited conditions. Microbiol Res. 2022;252:126858.
- Kumar R, Meena VS, Rajput VD. Mechanisms of phosphate solubilization by rhizobacteria: A case study in tomato. Arch Microbiol. 2024;206:101-115.
- Kumar S, Stecher G, Tamura K. MEGA11: Molecular evolutionary genetics analysis version 11.0 for bigger datasets. Mol Biol Evol. 2023;40(5):1890-1894.
- Larran S, Santamarina Siurana MP, Roselló Caselles J, Simón MR, Perelló A. In vitro antagonistic activity of Trichoderma harzianum against Fusarium sudanense causing seedling blight and seed rot on wheat. ACS Omega. 2020;5(36):23276-23283.
CrossRef - Mandal S, Roy A, Banerjee R. Soil enrichment and pathogen suppression through PGPR consortia in tomato crops. Biocatal Agric Biotechnol. 2024;54:102903.
- Meena KK, Sorty AM, Bitla UM. Phosphate solubilizing microbes: Sustainable approach for managing phosphorus deficiency in agricultural soils. Springer Sci Rev. 2020;5(2):1-19.
- Meena VS, Kumar S, Sharma P. Bioefficacy of PGPR in alkaline and saline tomato fields. Agron Sustain Dev. 2024;44:14.
- Naveed M, Qureshi MA, Zahir ZA, Hussain MB. Perspectives of PGPR-mediated growth enhancement in stress-prone ecosystems. Appl Soil Ecol. 2020;147:103367.
- Nema RK, Sharma L, Giri S. Phosphate-solubilizing rhizobacteria: Multifunctional traits for tomato growth. Rhizosphere. 2024;25:100662.
- Pandey A, Singh B, Kumar P, Maheshwari DK. Phosphate solubilizing microorganisms and their role in sustainable agriculture. World J Microbiol Biotechnol. 2021;37(8):1-13.
- Patel H, Singh N, Chauhan R. Siderophore-mediated iron uptake in tomato by PGPR. Environ Microbiol Rep. 2023;15(6):680-692.
- Rana A, Chauhan D, Bharti M. Metagenomic insights into PGPR genes from tomato rhizosphere. Sci Rep. 2023;13:7712.
- Rathi M, Gaur A. Phosphate solubilizing microorganisms: Recent advancements and their role in sustainable agriculture. Soil Biol Biochem. 2022;163:108449.
- Roy B, Maitra D, Biswas A. Efficacy of high-altitude biofilm-forming novel Bacillus subtilis species as plant growth-promoting rhizobacteria on Zea mays L. Appl Biochem Biotechnol. 2024;196(2):643-666.
CrossRef - Schwyn B, Neilands J. Universal chemical assay for the detection and determination of siderophores. Anal Biochem. 1987;160(1):47-56.
CrossRef - Sharma P, Bhatt P. Quantification of siderophores and phosphate solubilization in native PGPR. J Environ Biol. 2025;46(1):88-95.
- Sharma S, Mathur P, Goswami M, Dey N. Phosphate solubilizing bacteria in agriculture: Mechanism and diversity of solubilization. Int J Curr Microbiol Appl Sci. 2020;9(9):2281-2295.
- Shivay YS, Singh RK, Chaudhary R. Microbial inoculants in nutrient-deficient soils: Role of rhizobacteria. Front Microbiol. 2023;14:1112150.
- Singh D, Giri S, Patel DK. Rhizospheric microbes in plant health and nutrient cycling. Appl Soil Ecol. 2023;191:104965.
- Sinha R, Dubey A, Saxena A. Integrated microbial strategies to improve soil fertility in tomato fields. J Soil Sci Plant Nutr. 2025;25(1):45-58.
- Sun Y, Wu J, Shang X. Screening of siderophore-producing bacteria and their effects on promoting the growth of plants. Curr Microbiol. 2022;79(5):150.
CrossRef - Thakur P, Bhattacharya D, Singh A. Isolation of phosphate-solubilizing rhizobacteria and evaluation of their plant growth traits. Environ Microbiome. 2024;19:34.
- Tripathi A, Mishra V, Tiwari R. Functional analysis of PGPR genes involved in nutrient solubilization. Microb Ecol. 2024;88(3):789-802.
- Ustiatik R, Nuraini Y, Handayanto E. Siderophore production of Hg-resistant endophytic bacteria isolated from local grass in Hg-contaminated soil. J Ecol Eng. 2021;22(5).
CrossRef - Yadav AN, Singh J, Rastegari AA, Gupta VK. Microbiome-based interventions for sustainable agriculture and ecosystem management. J Plant Growth Regul. 2022;41:1866-1885.
- Zhao F, Li Y, Wang S, Yang W, Zhao Y, Yang J. Isolation and characterization of phosphate solubilizing bacteria from rhizosphere soils of metal-contaminated plants. Environ Pollut. 2021;269: 116148.
- Zhao X, Yuan X, Xing Y. A meta-analysis on morphological, physiological and biochemical responses of plants with PGPR inoculation under drought stress. Plant Cell Environ. 2023;46(1):199-214.
CrossRef

